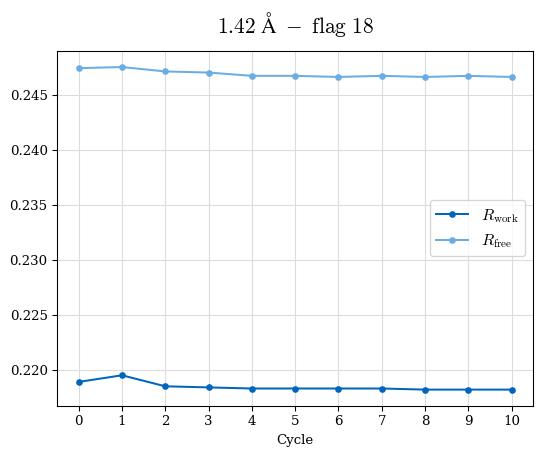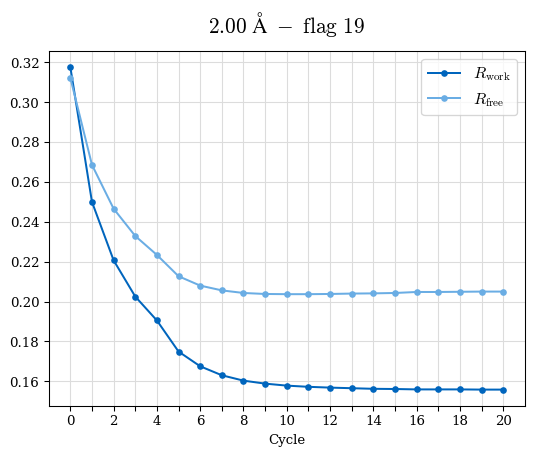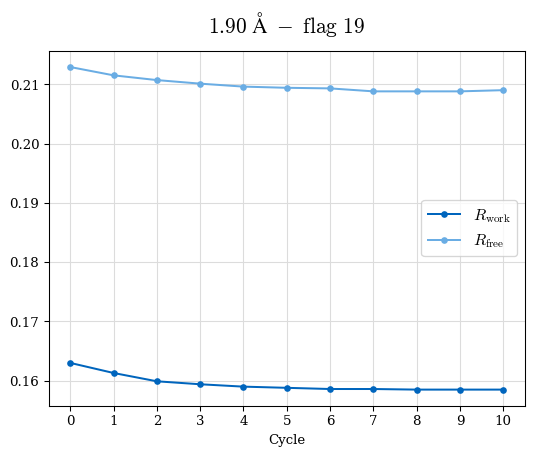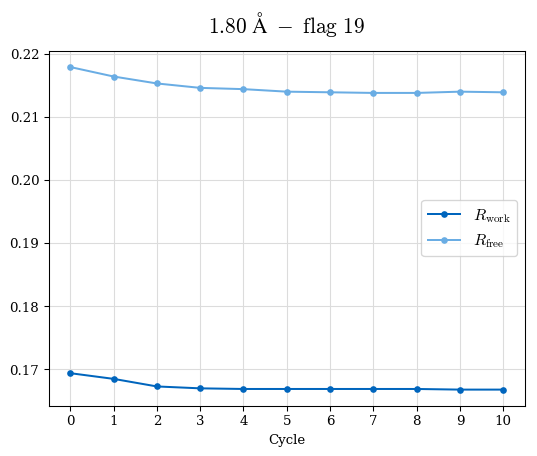
Raw data: CDO_complete_R-values.csv
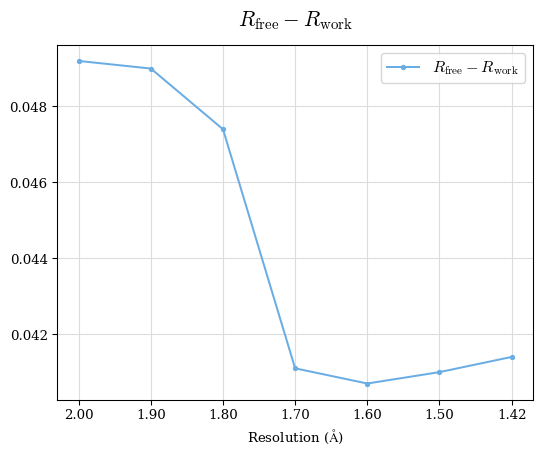
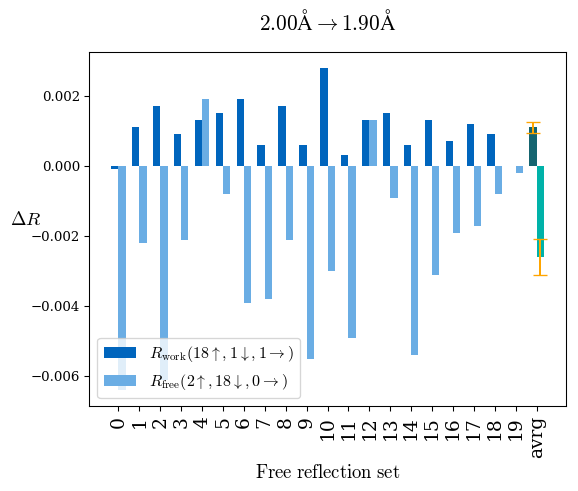
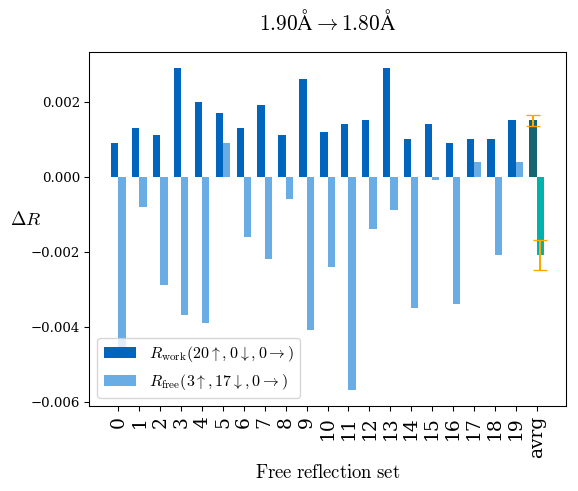
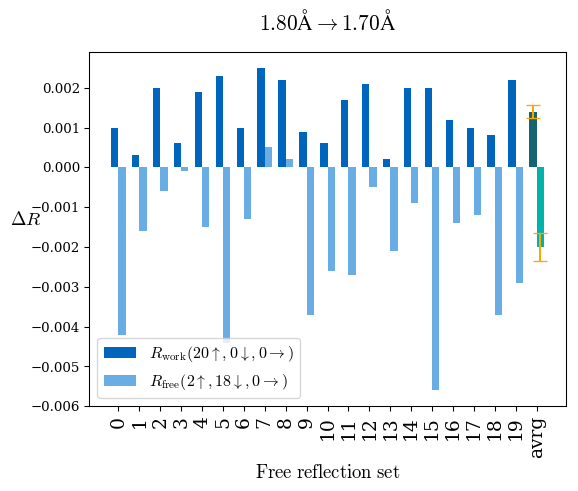
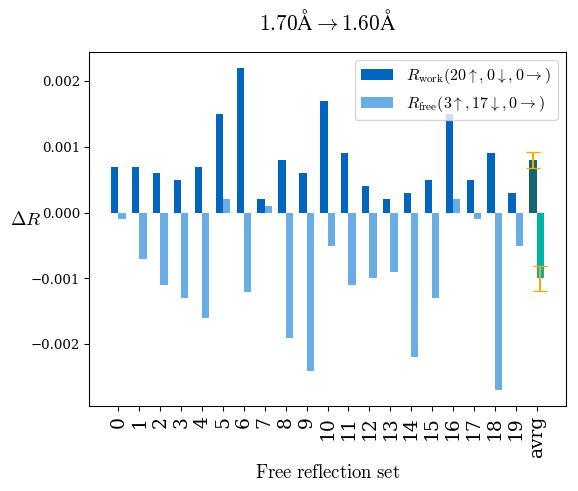
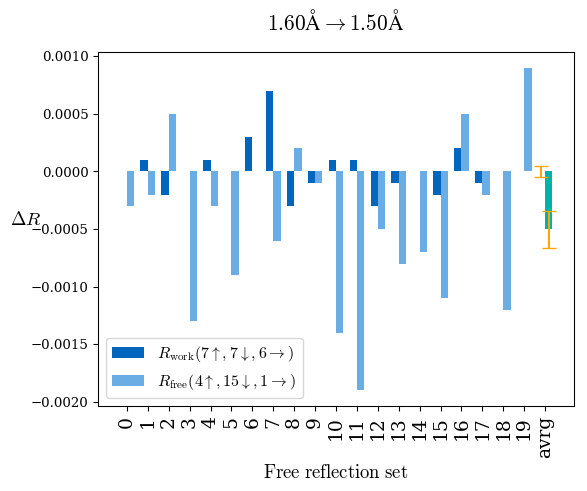
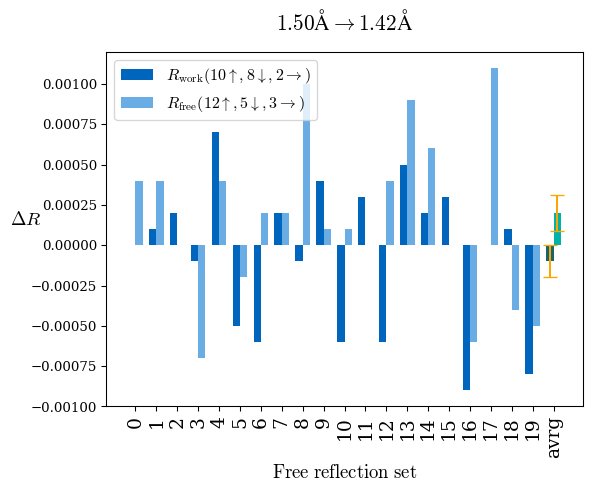
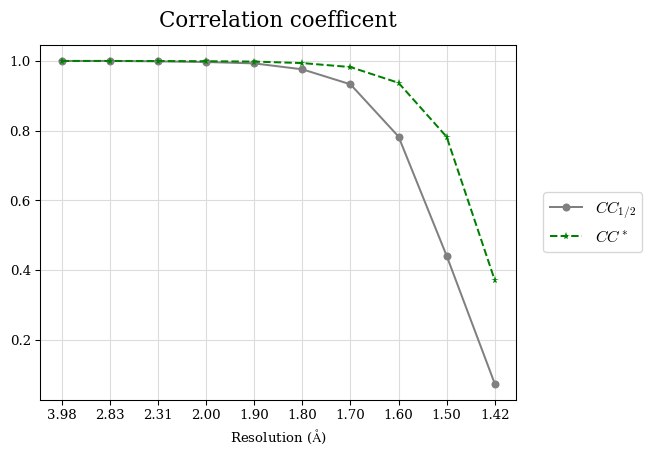
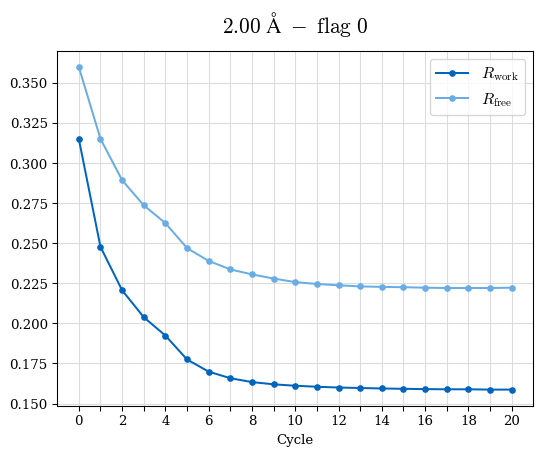
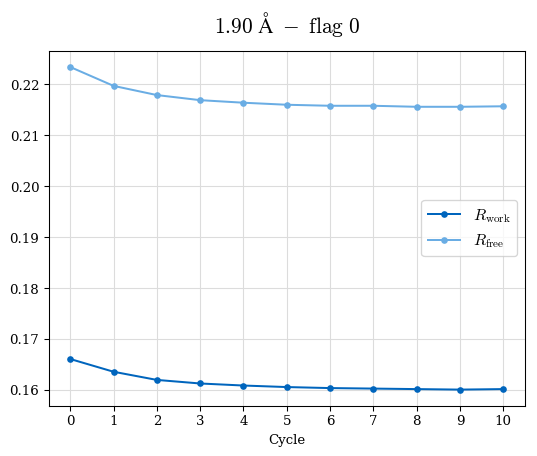
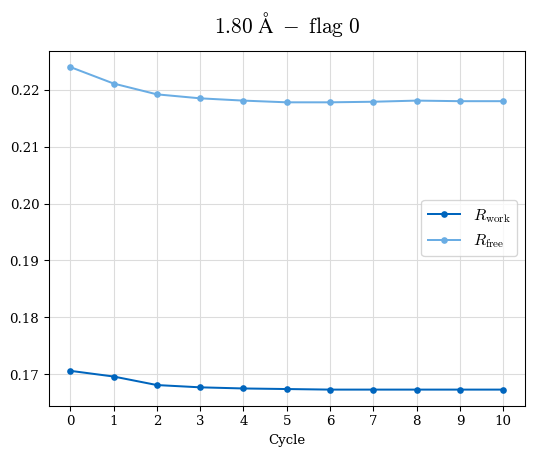
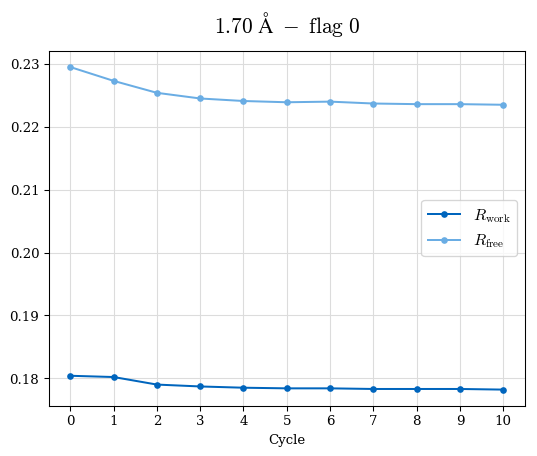
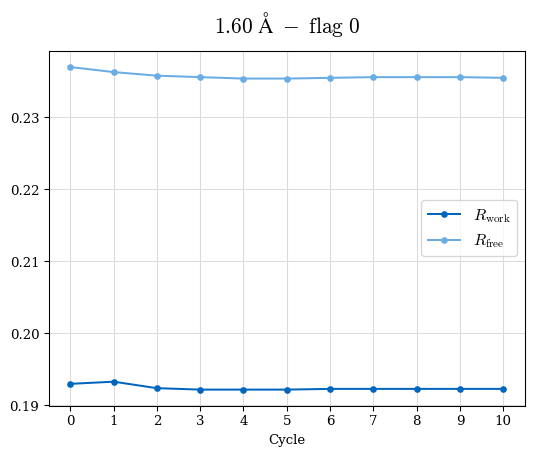
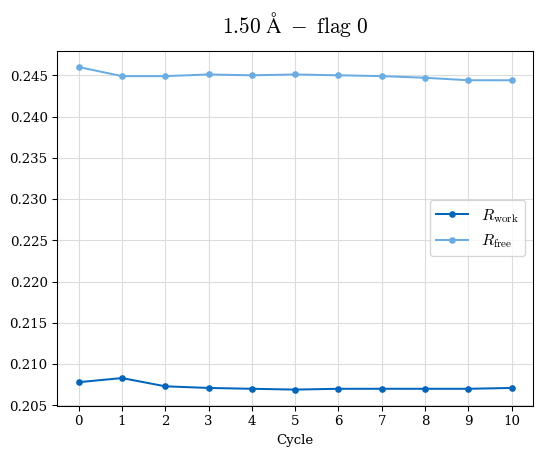
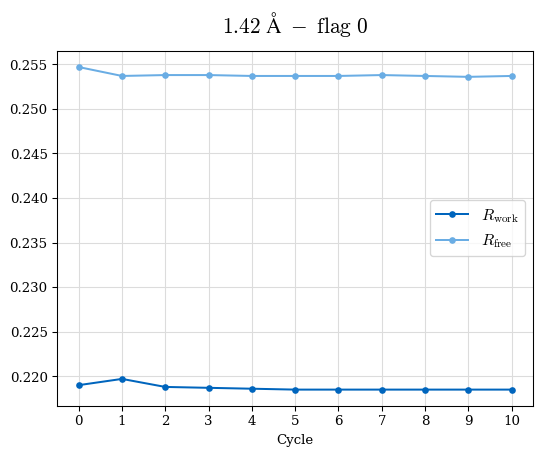
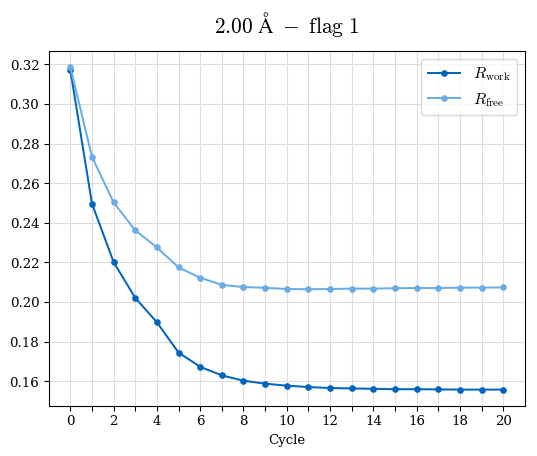
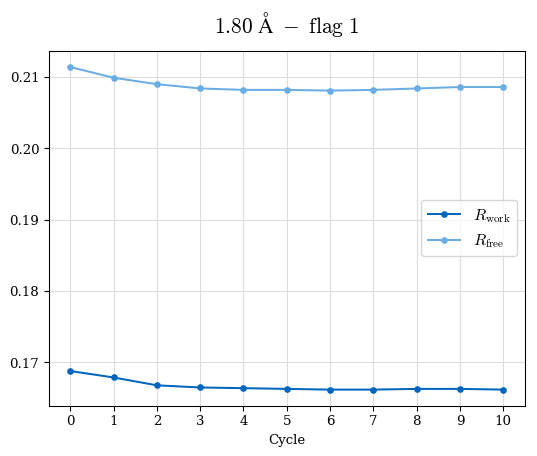
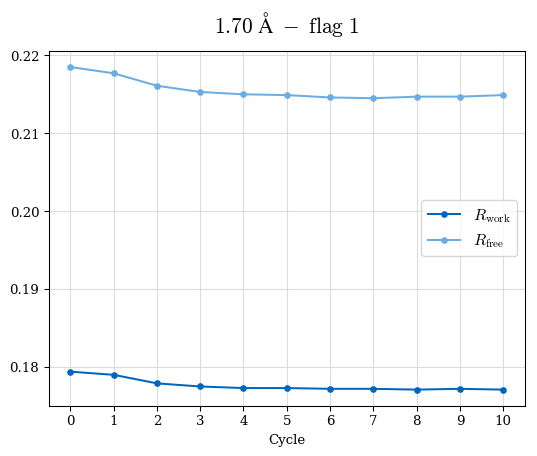
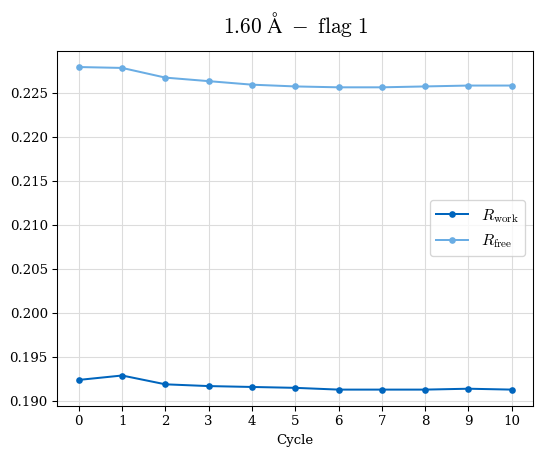
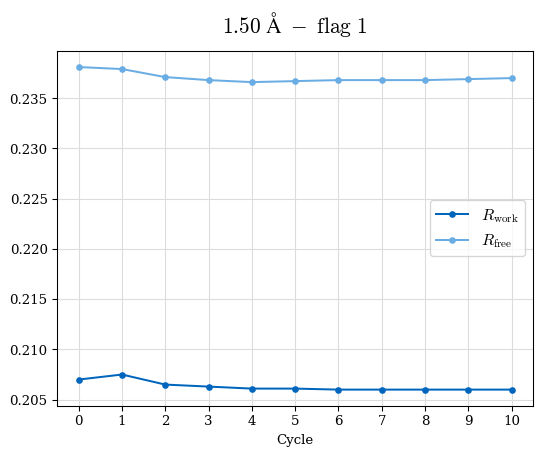
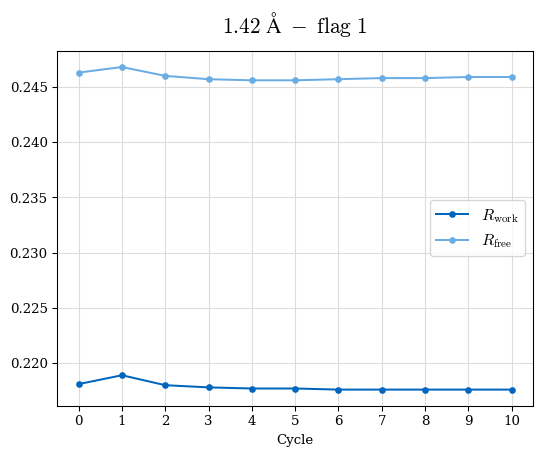
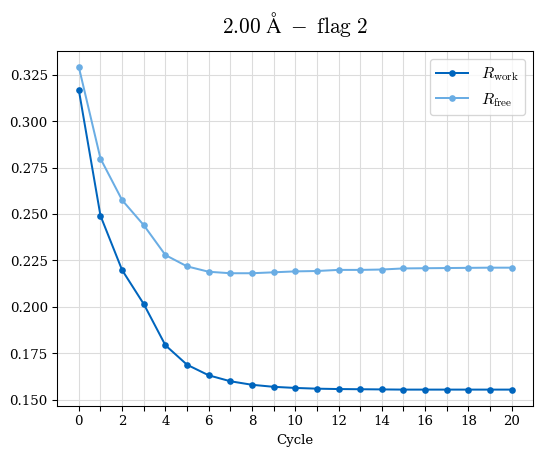
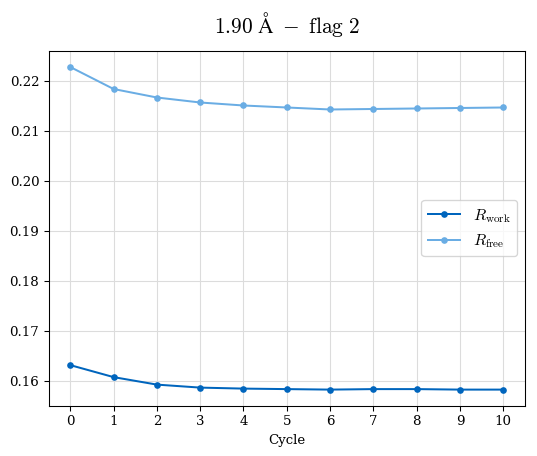
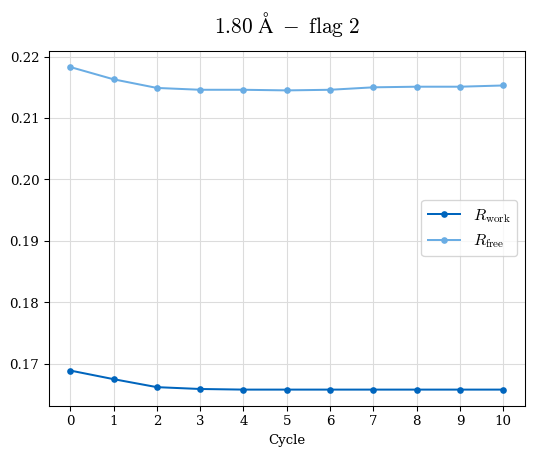
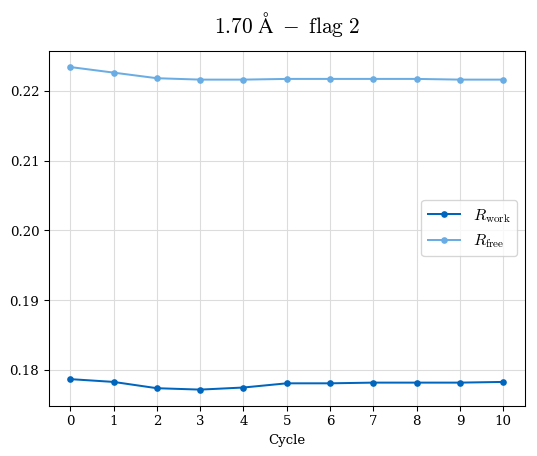
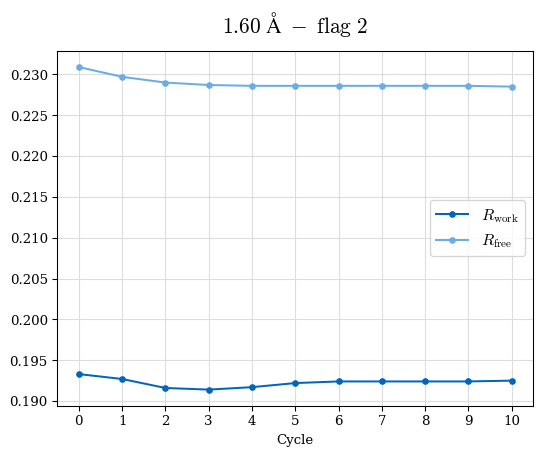
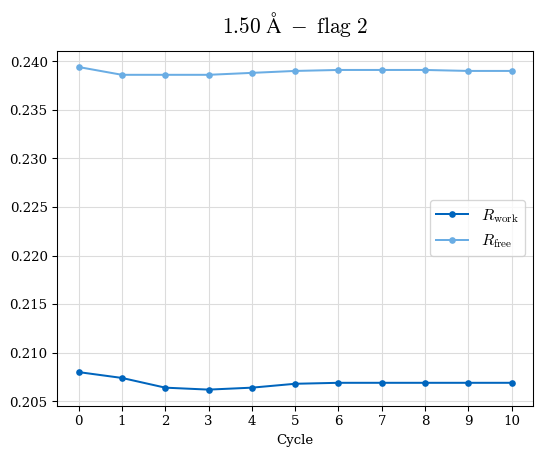
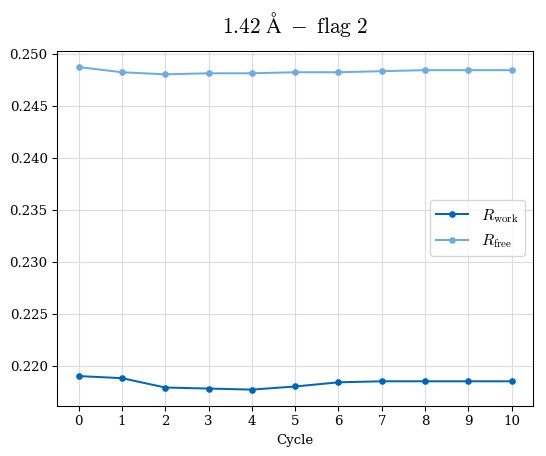
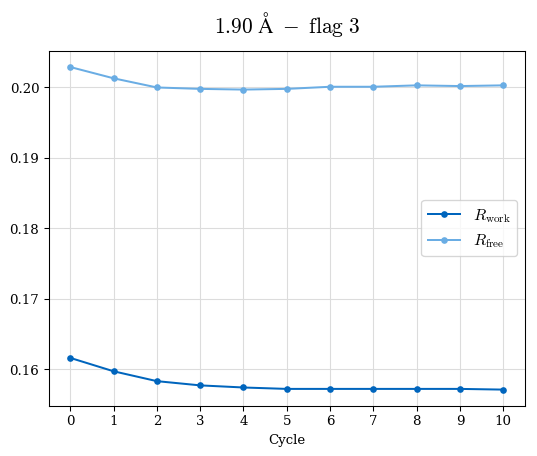
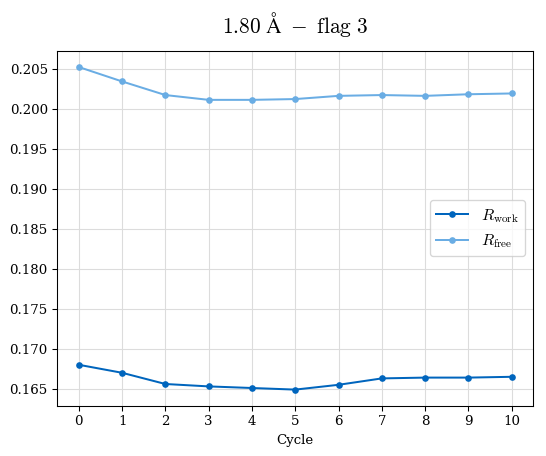
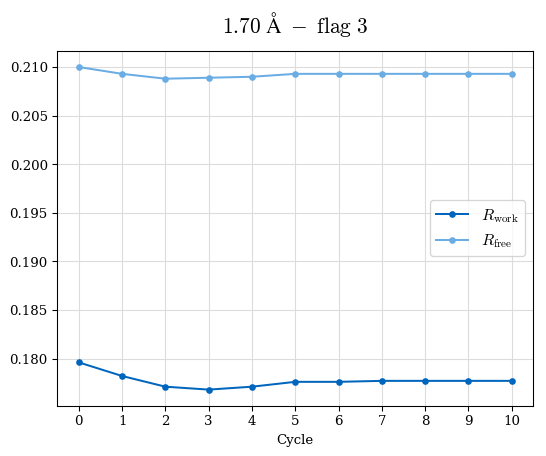
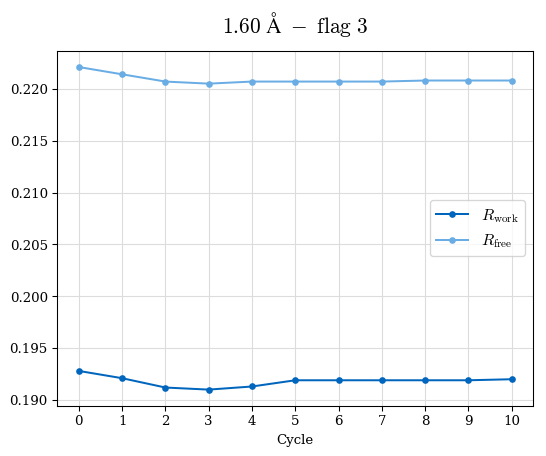
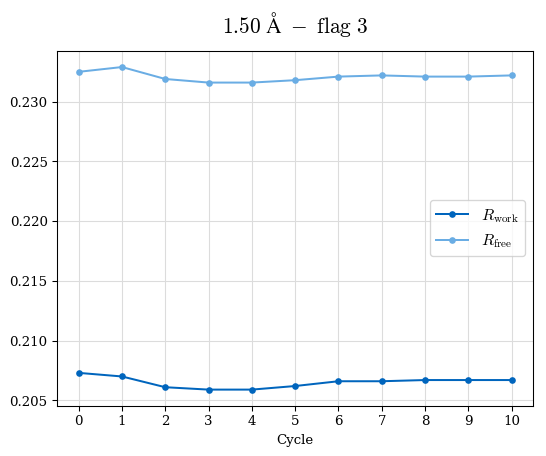
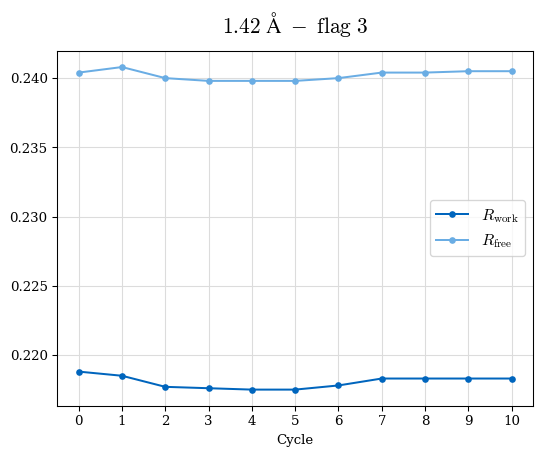
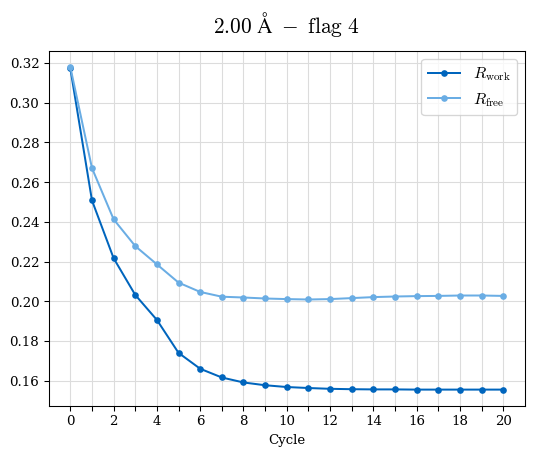
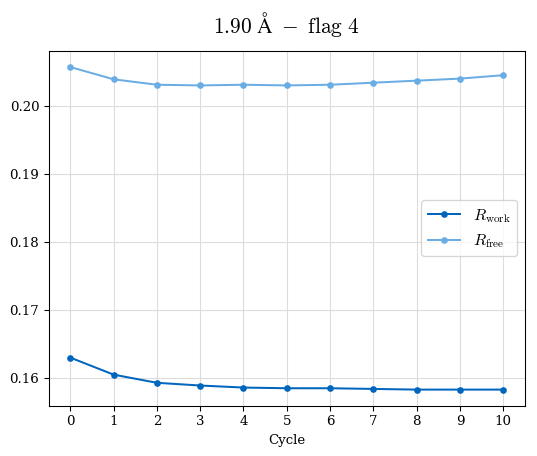
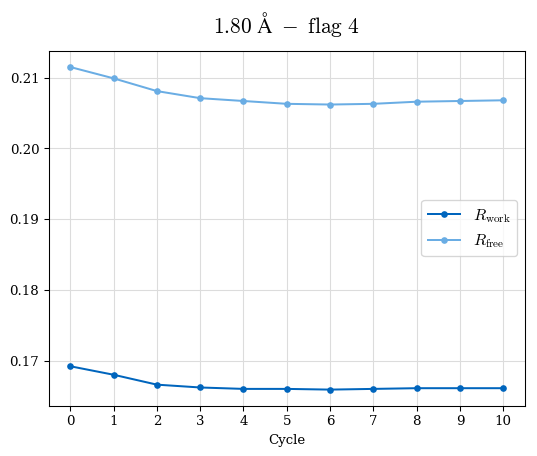
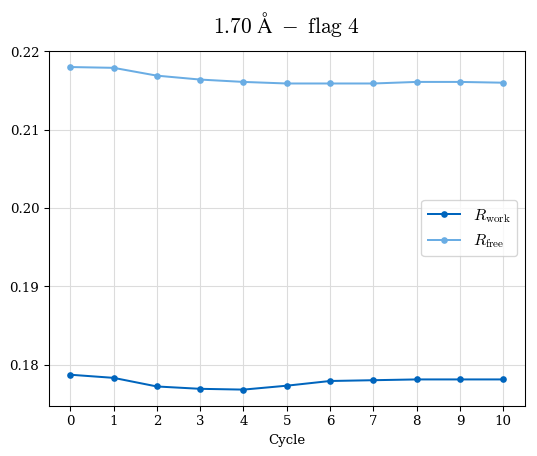
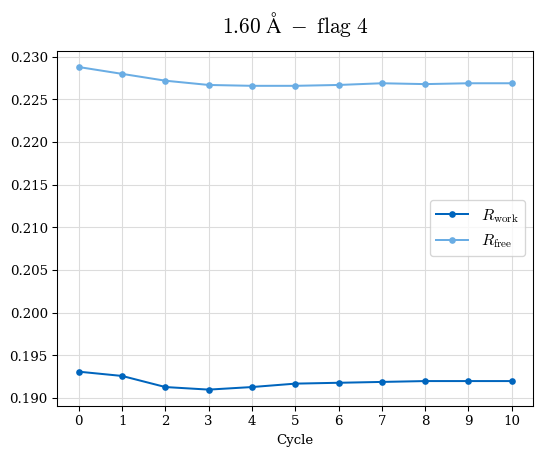
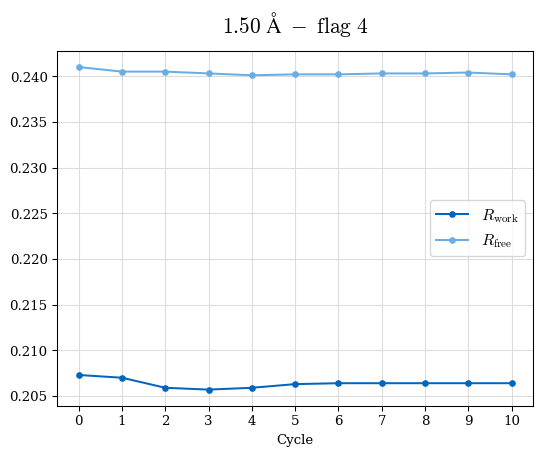
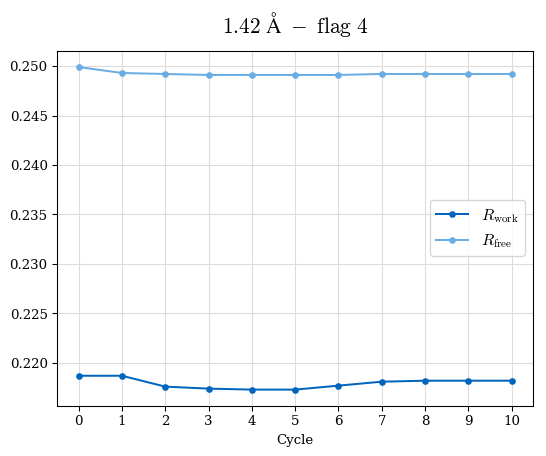
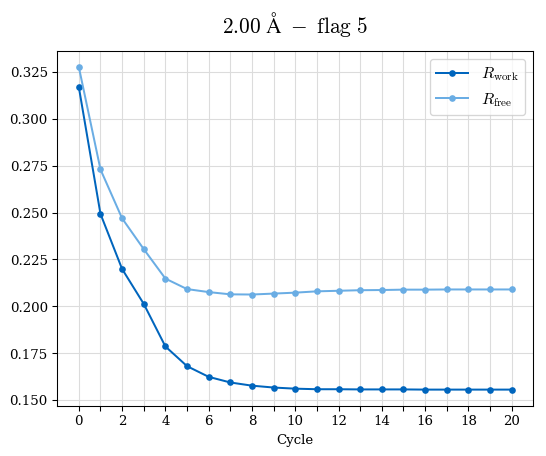
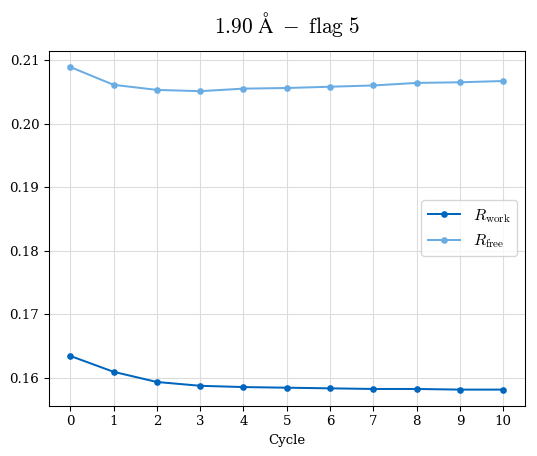
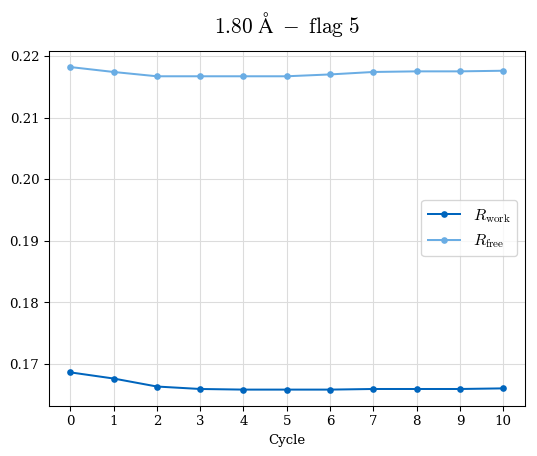
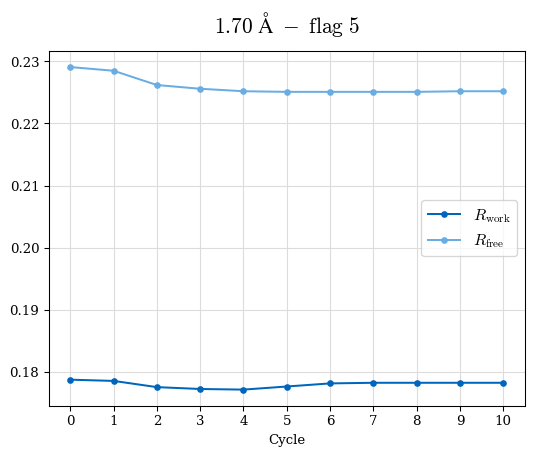
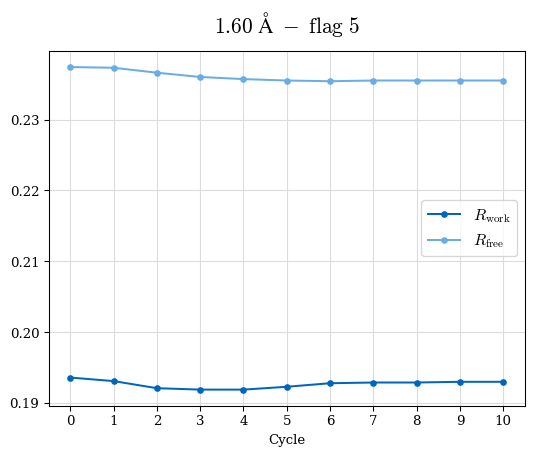
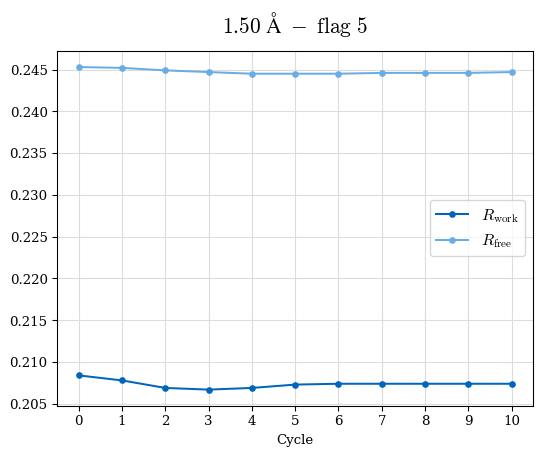
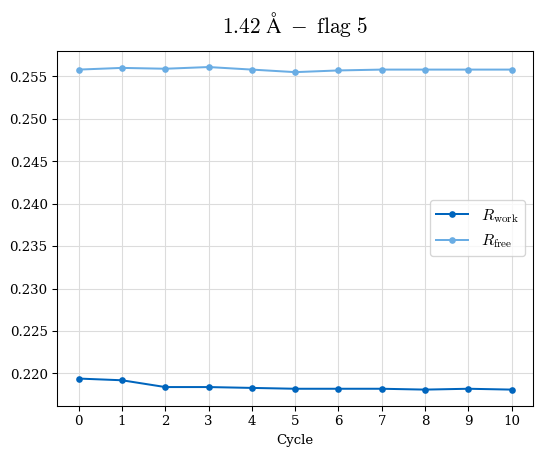
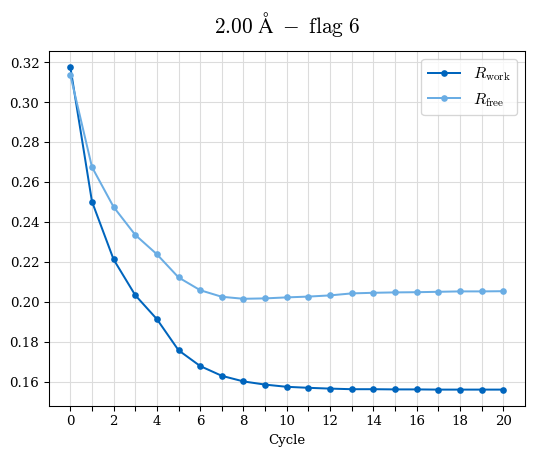
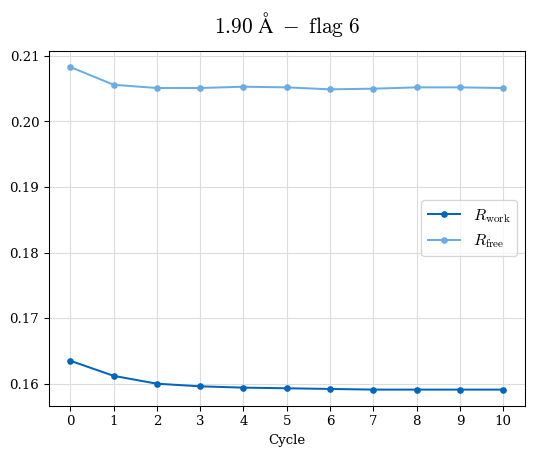
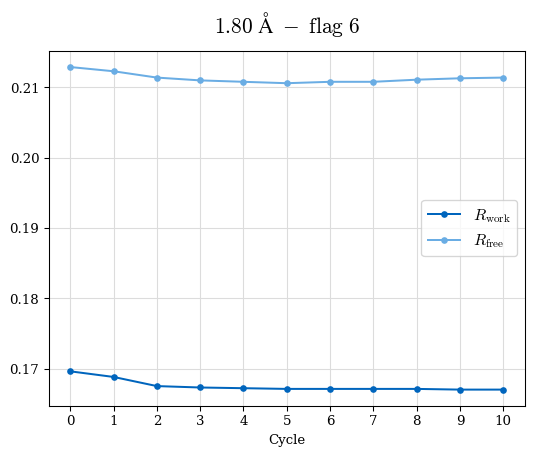
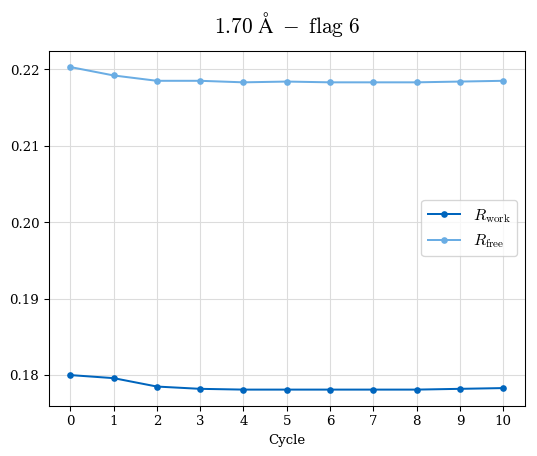
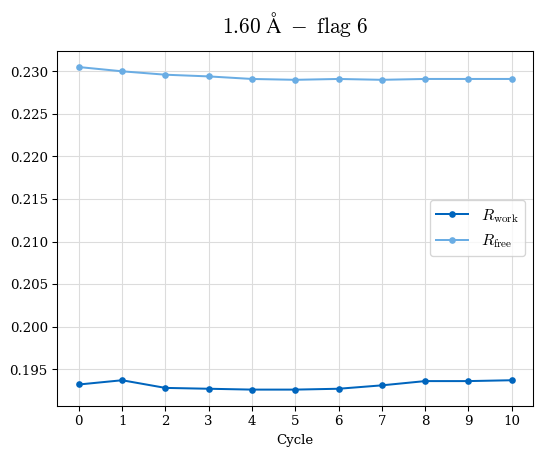
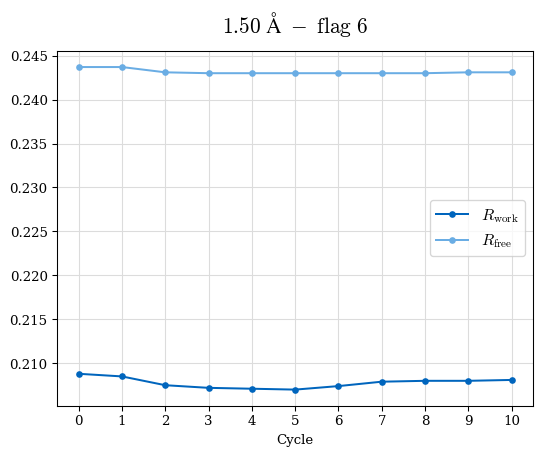
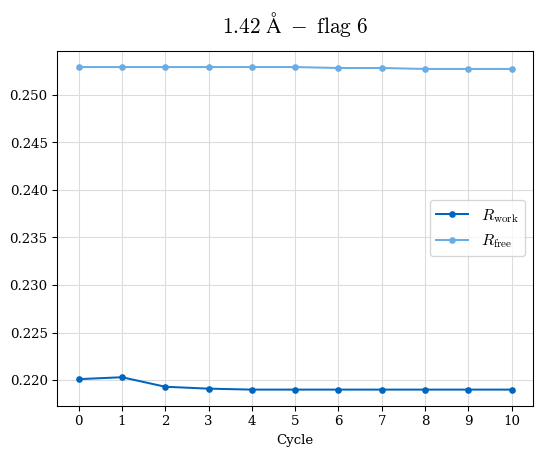
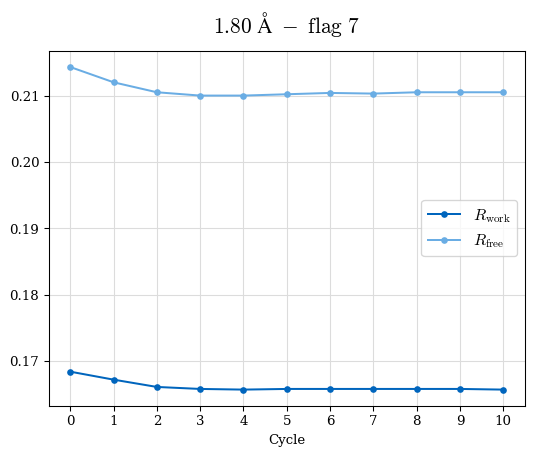
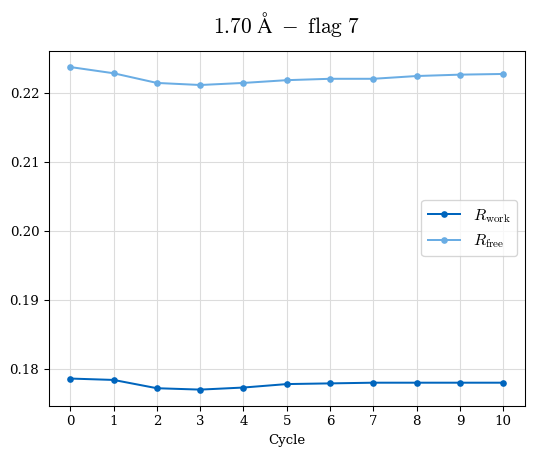
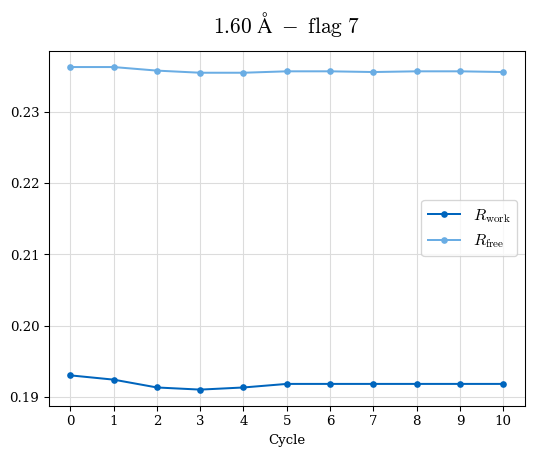
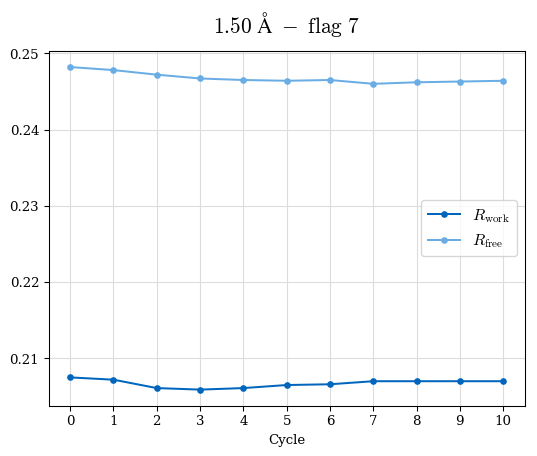
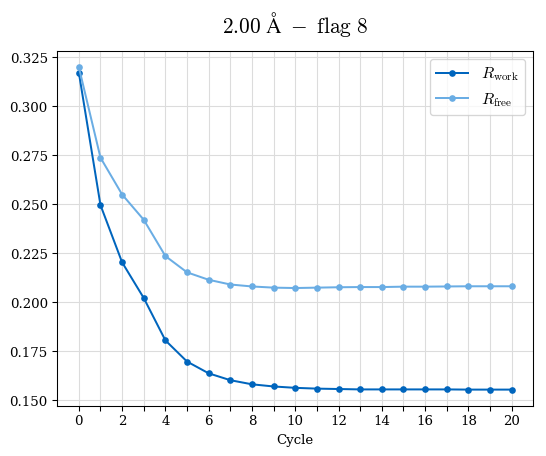
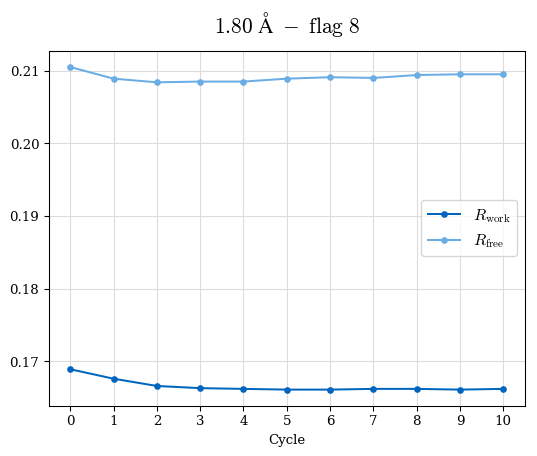
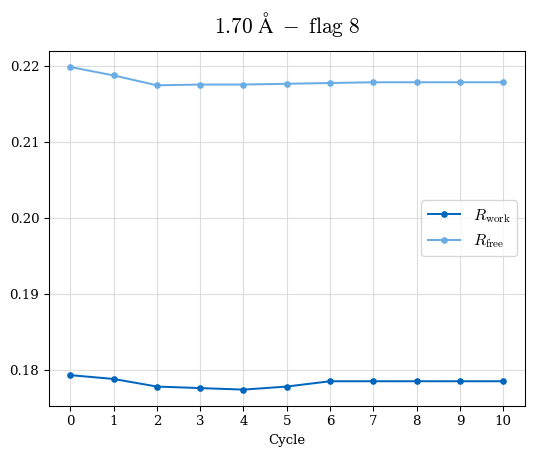
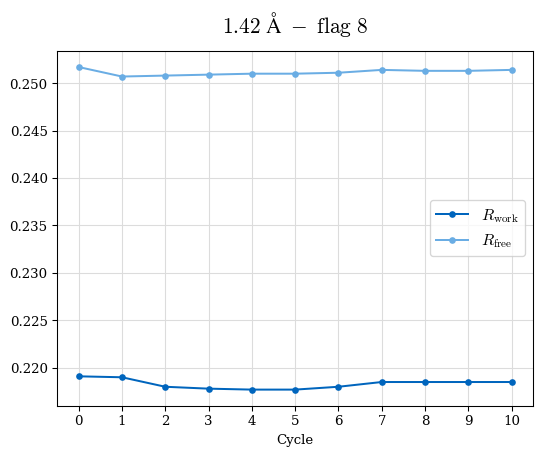
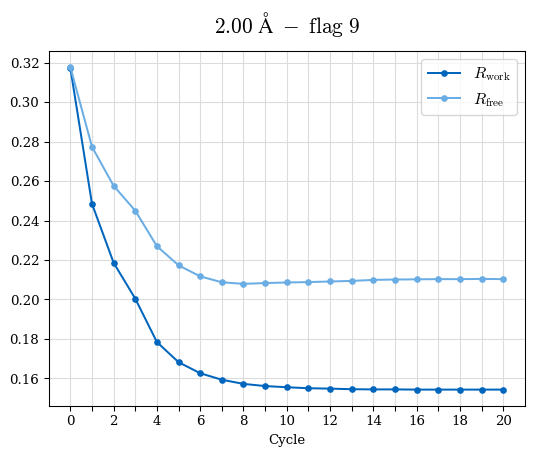
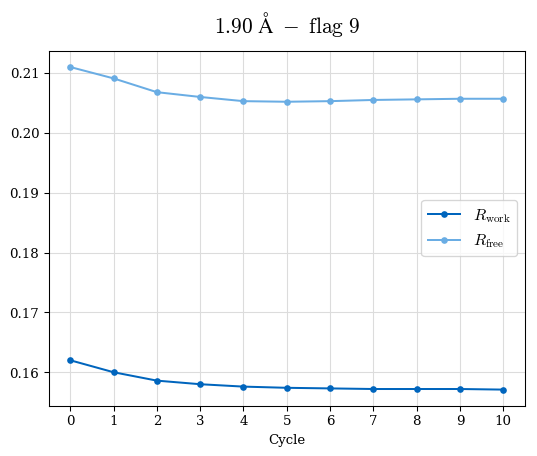
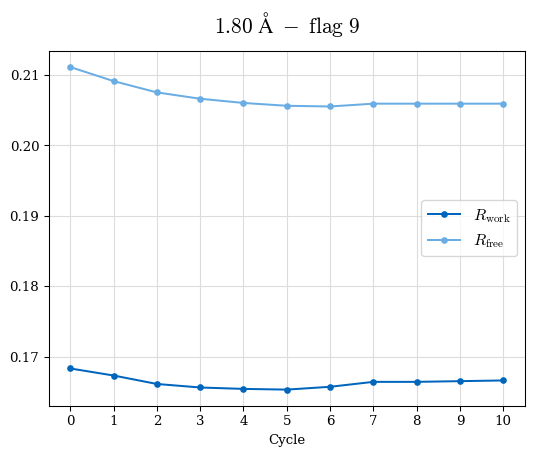
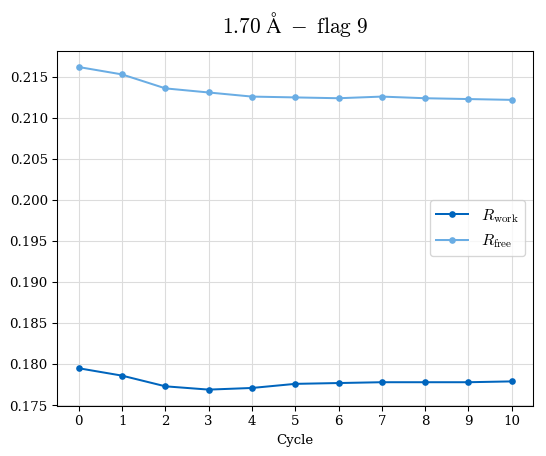
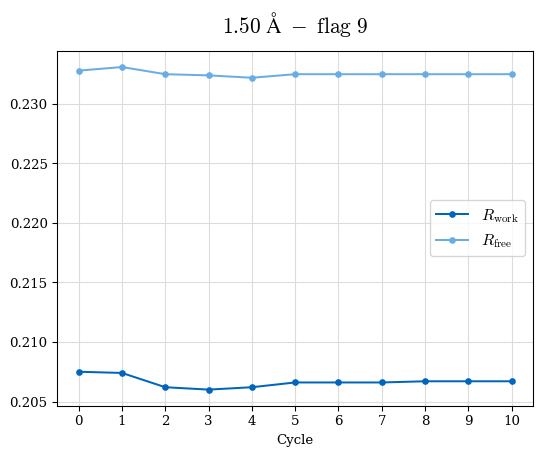
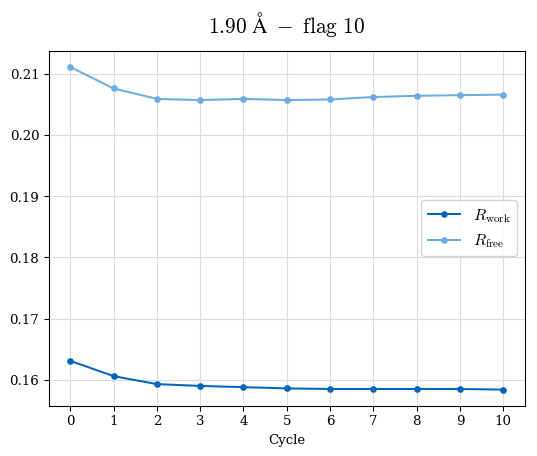
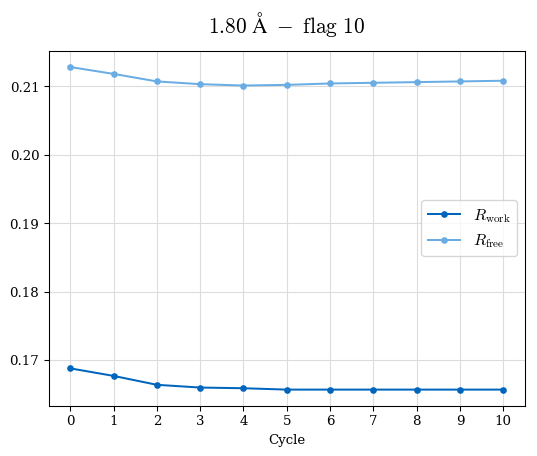
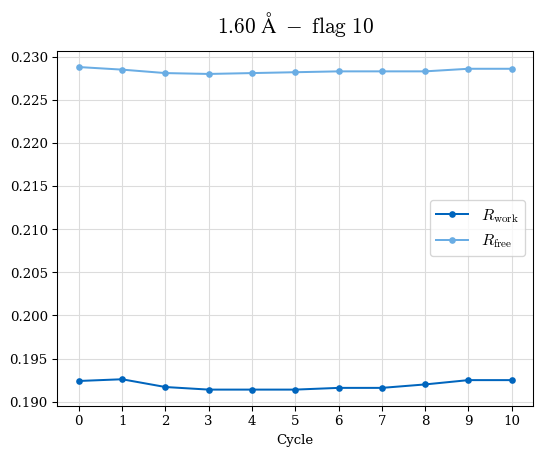
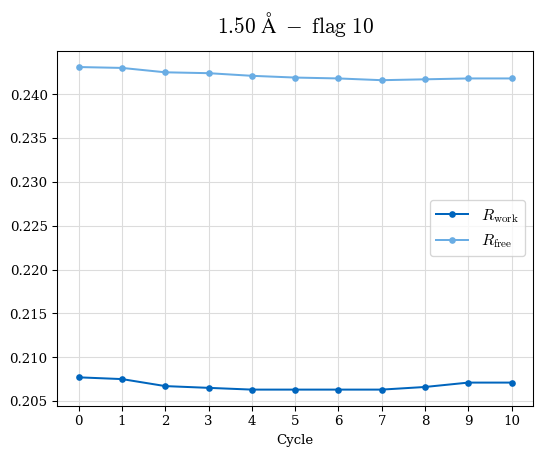
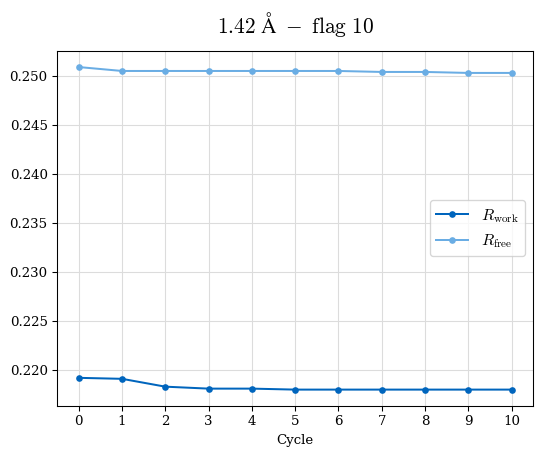
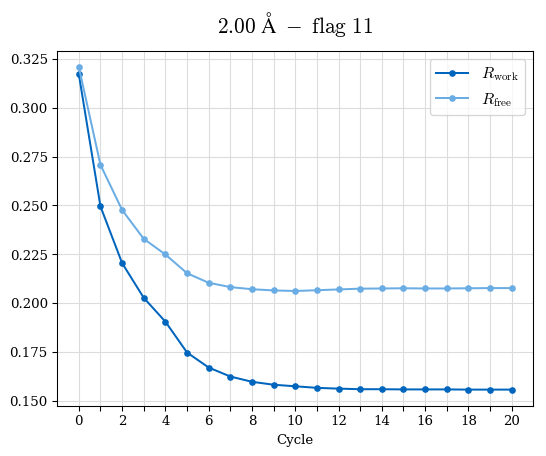
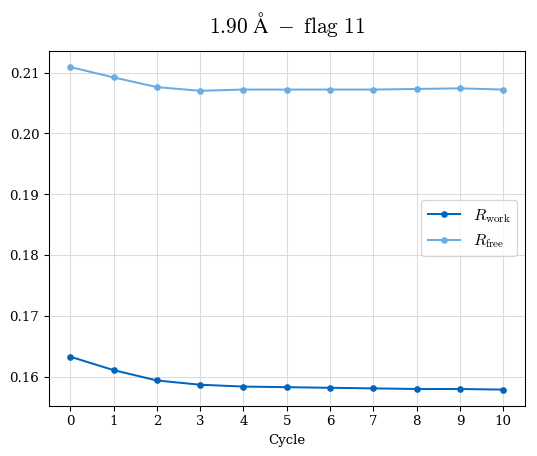
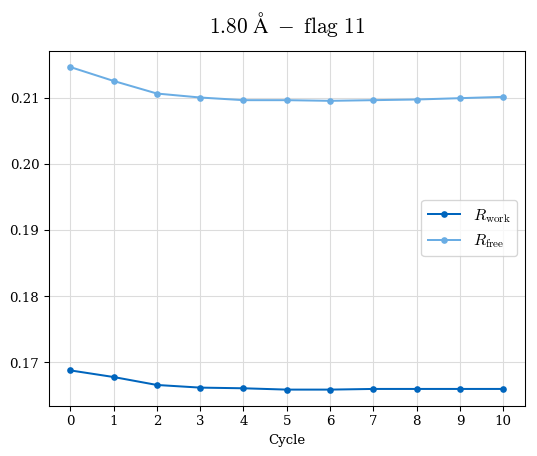
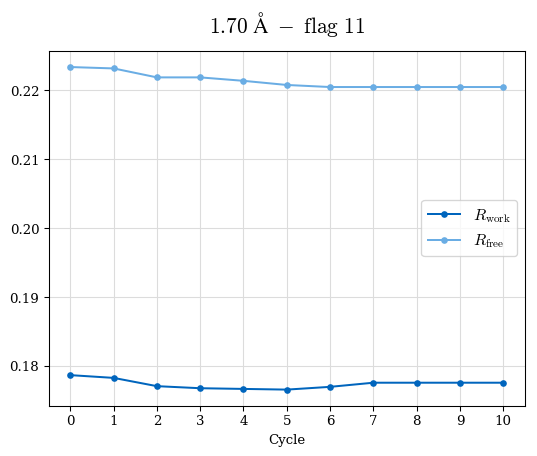
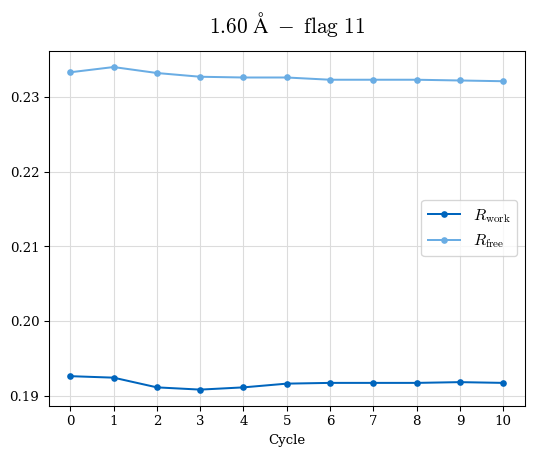
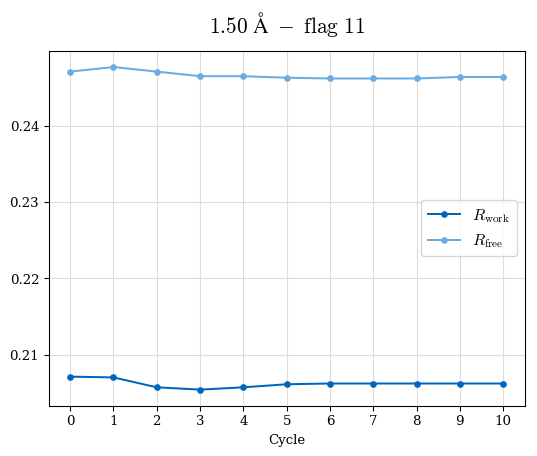
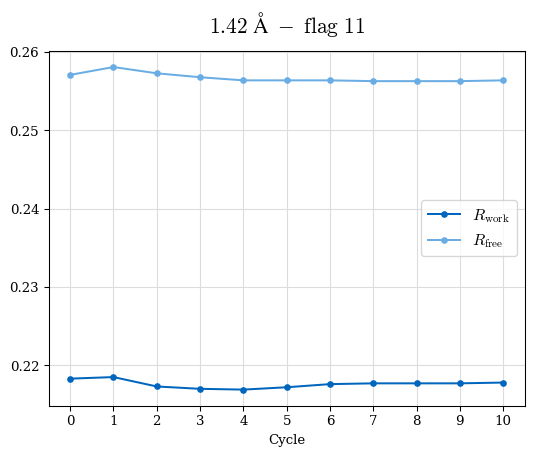
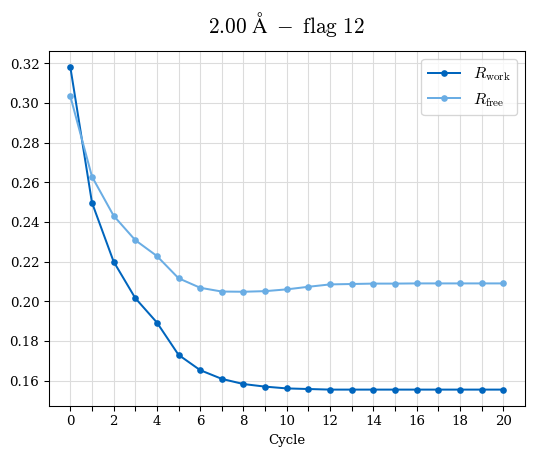
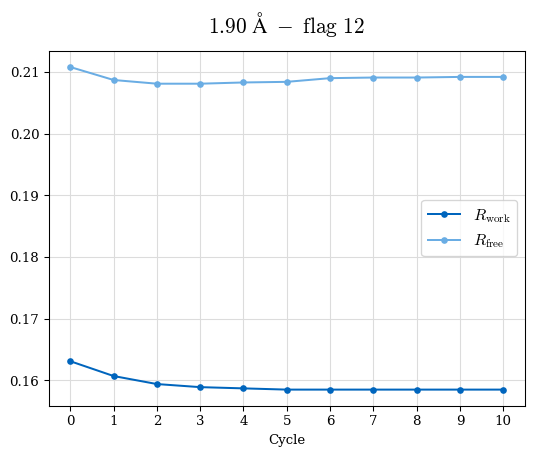
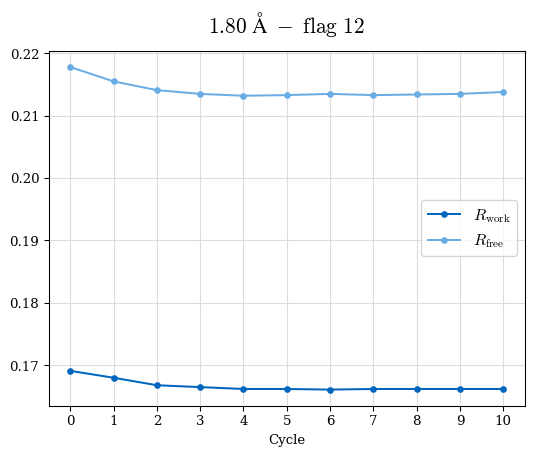
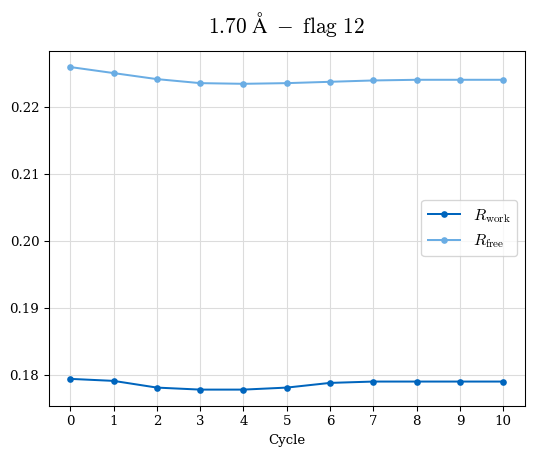
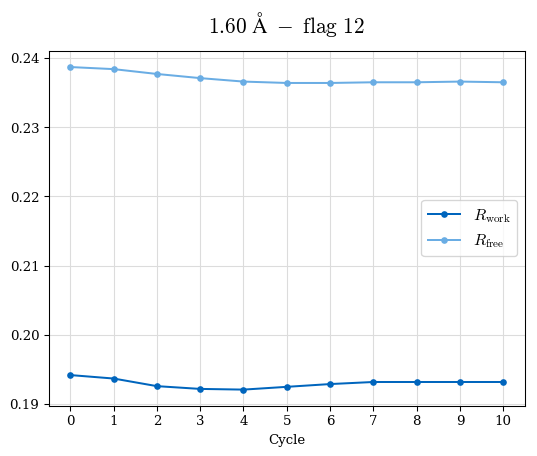
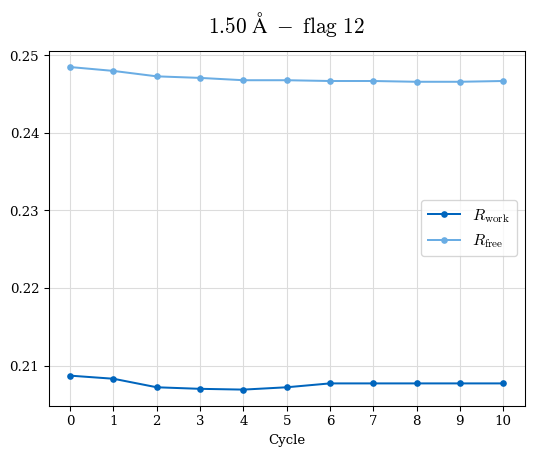
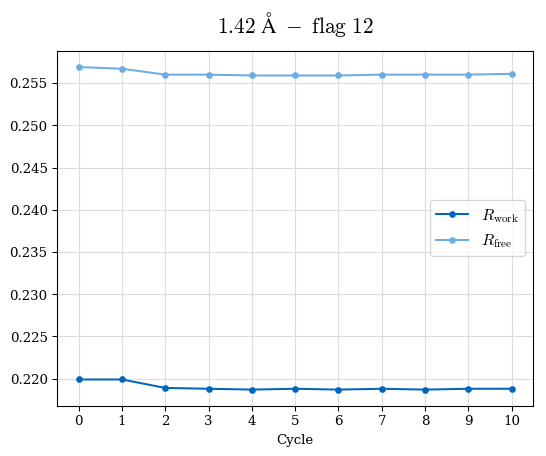
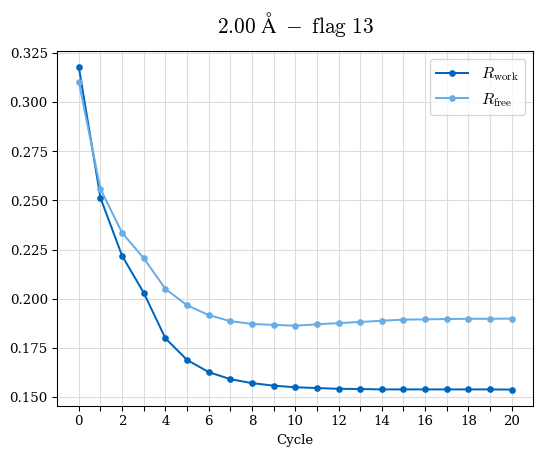
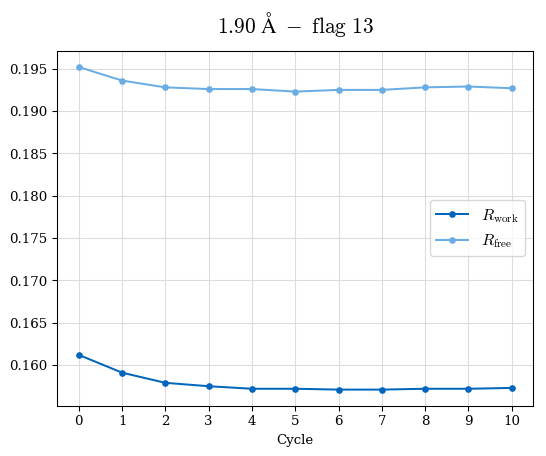
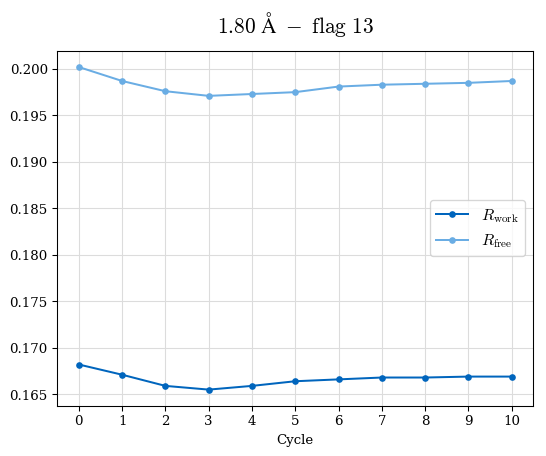
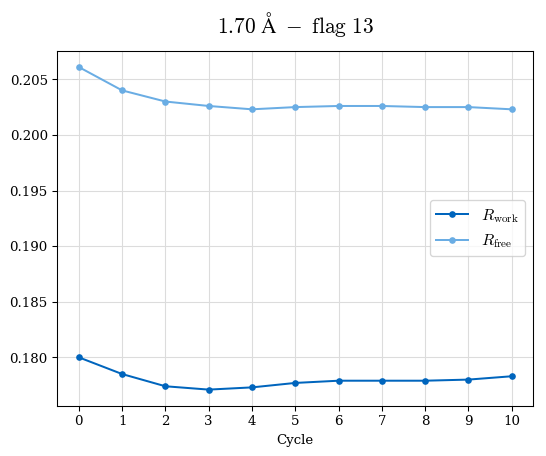
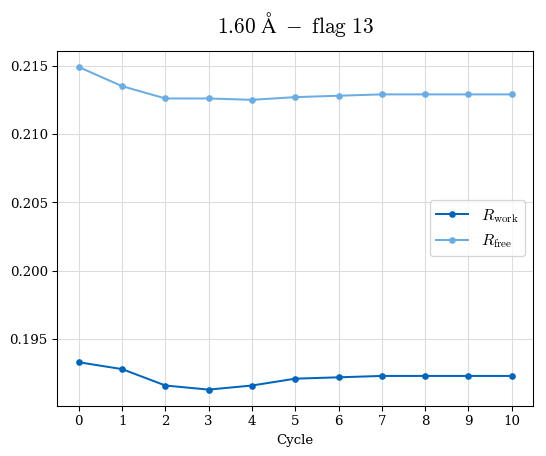
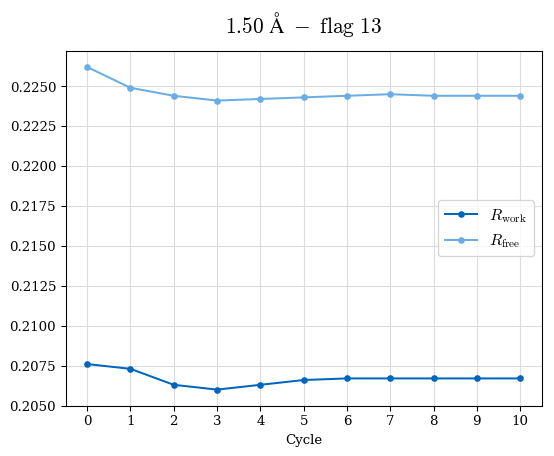
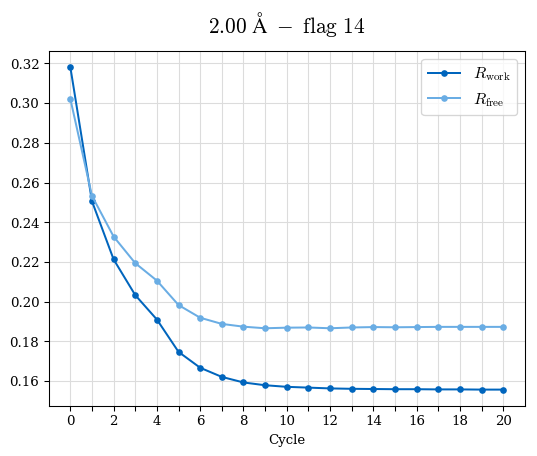
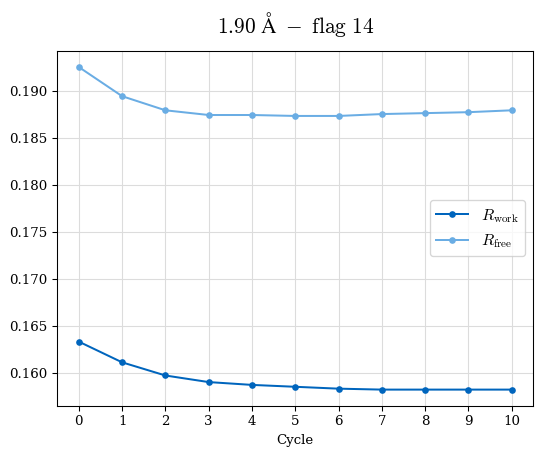
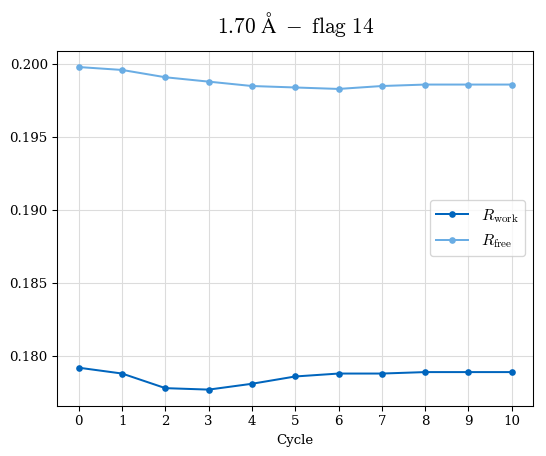
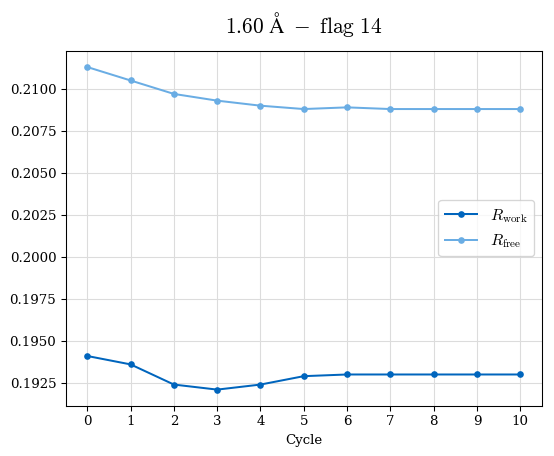
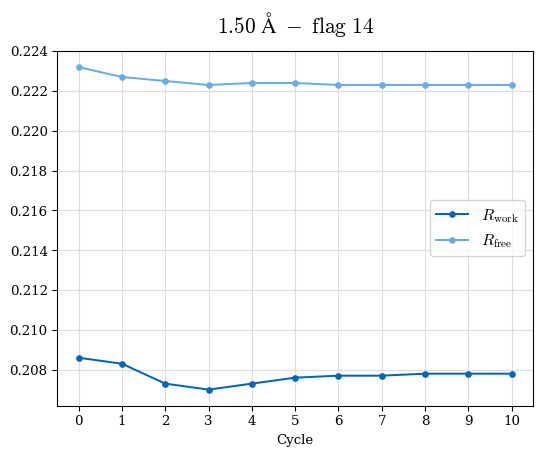
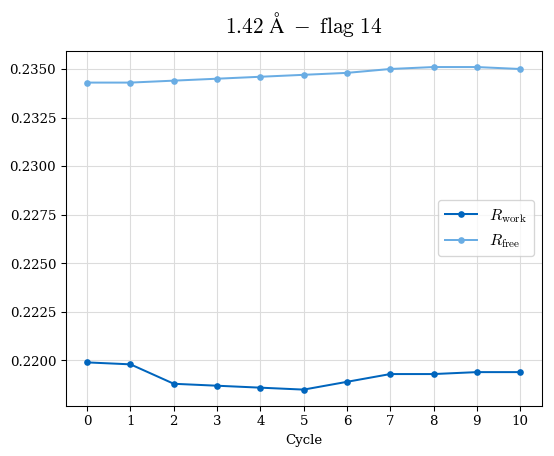
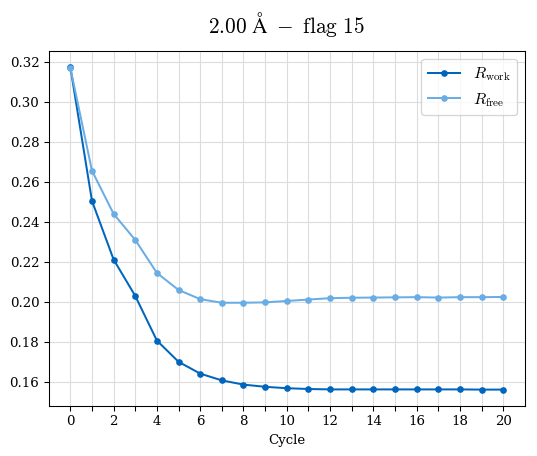
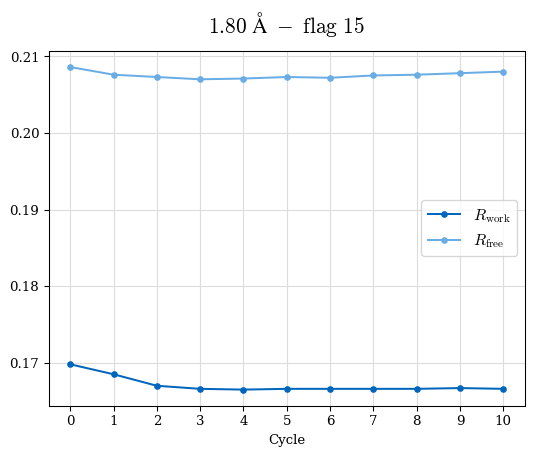
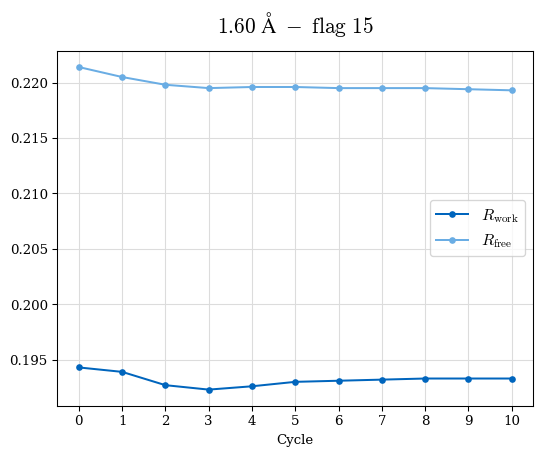
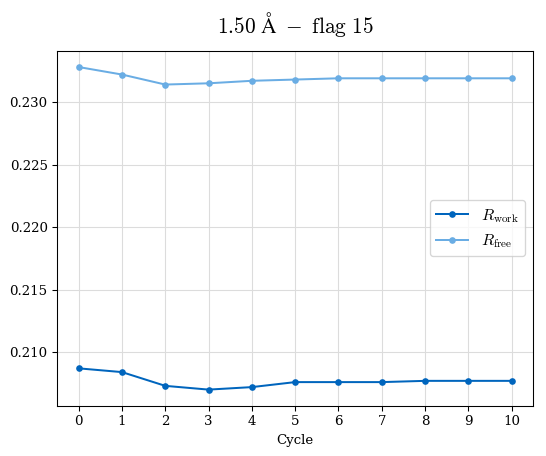
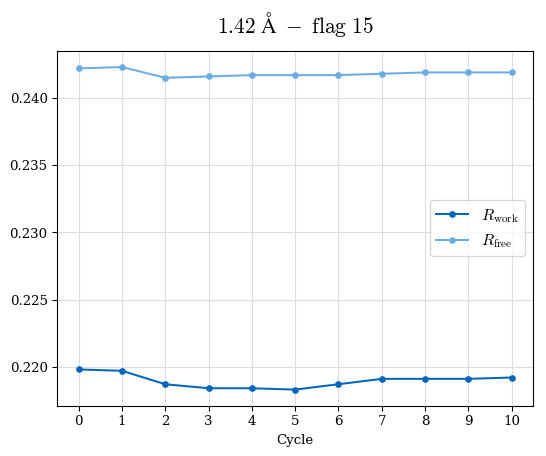
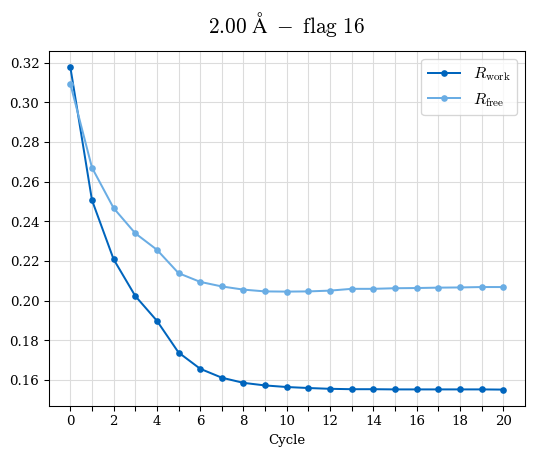
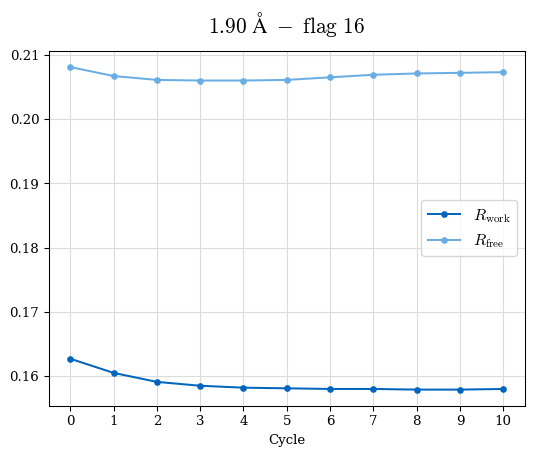
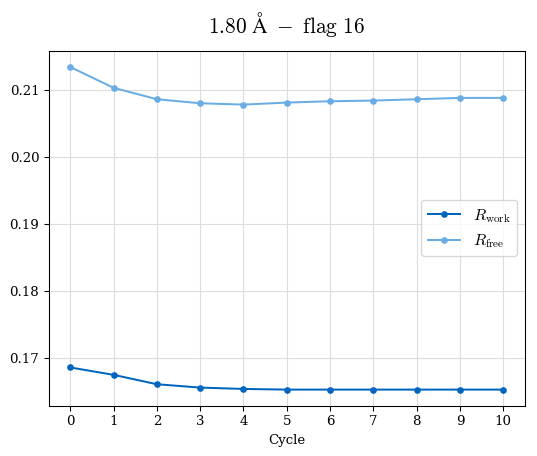
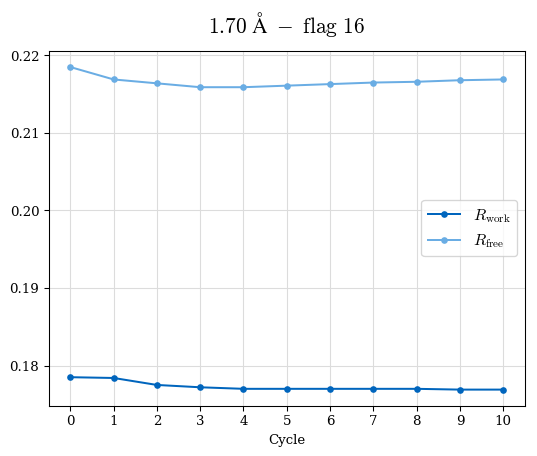
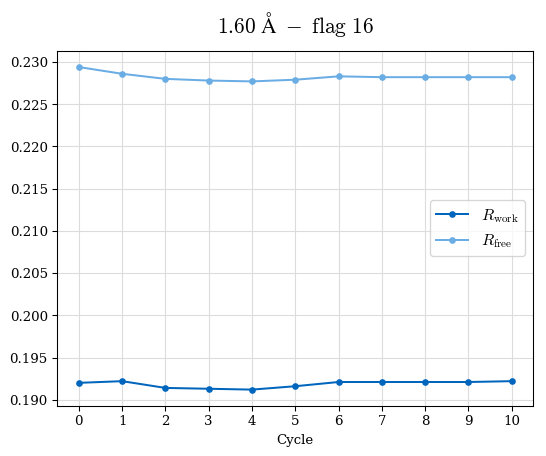
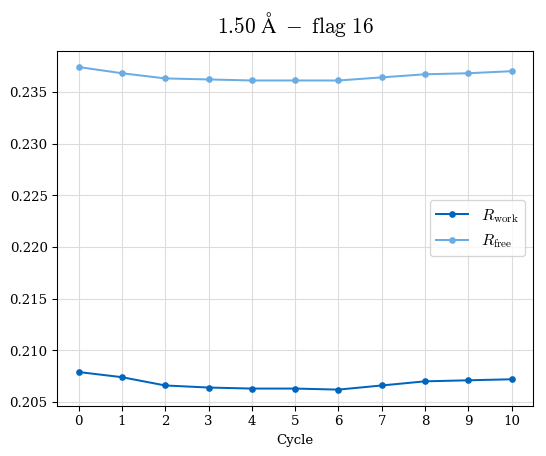
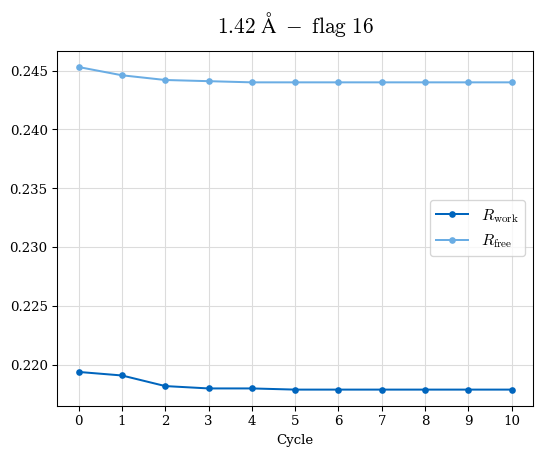
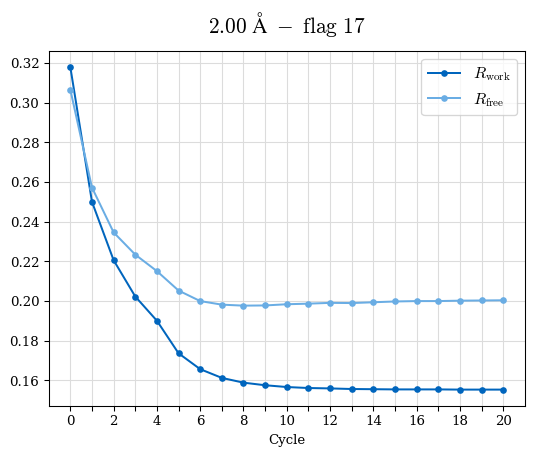
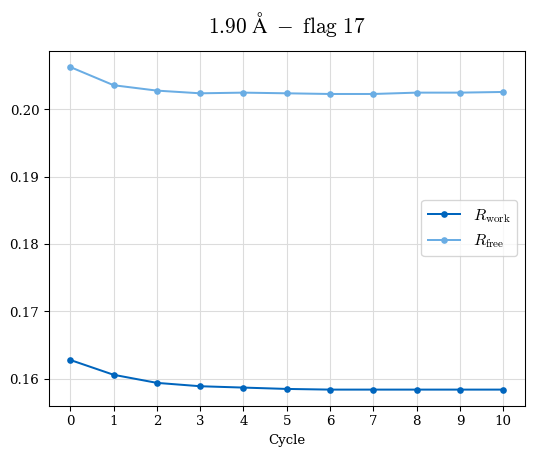
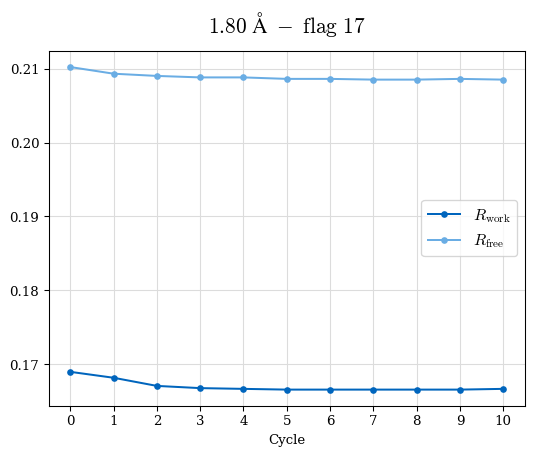
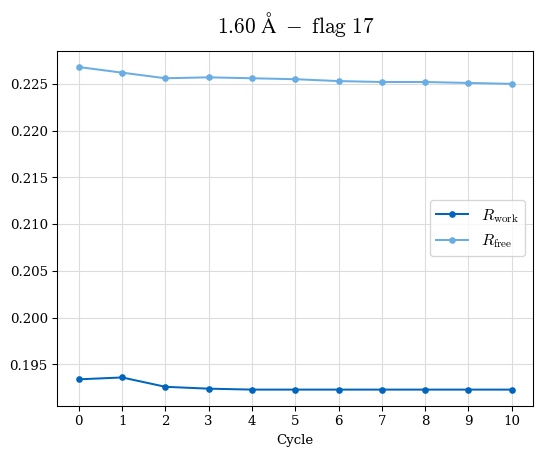
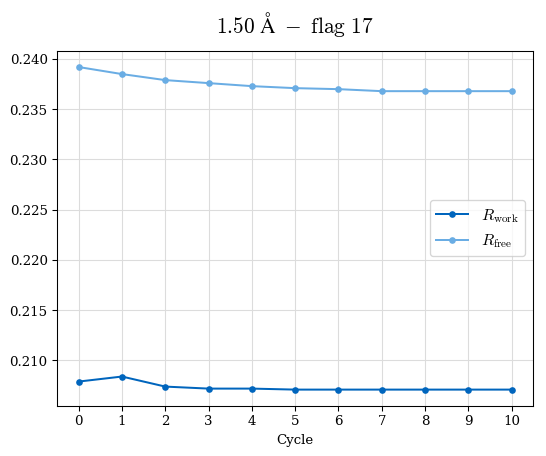
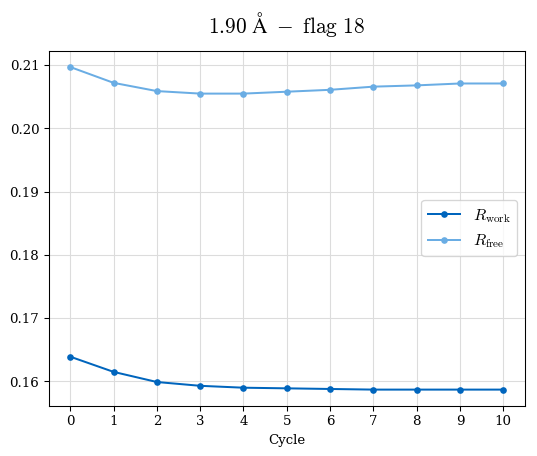
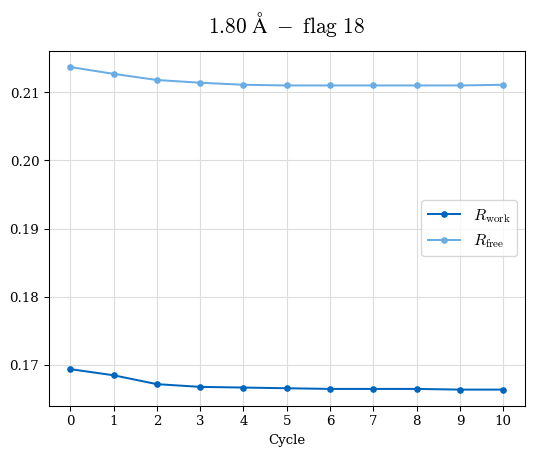
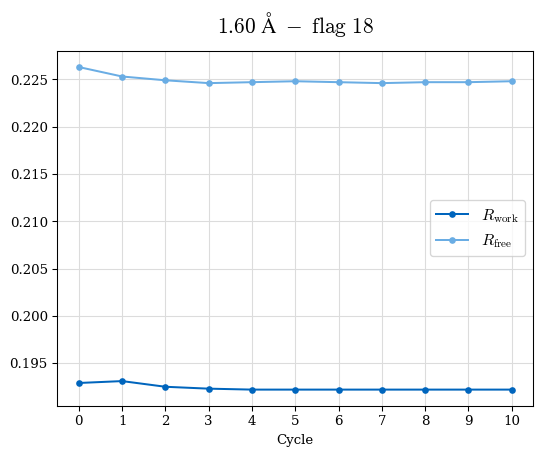
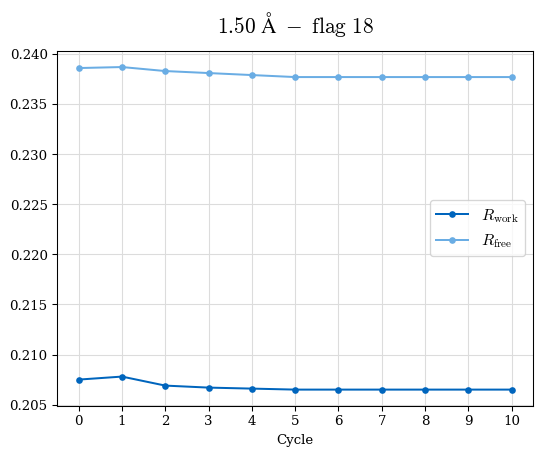
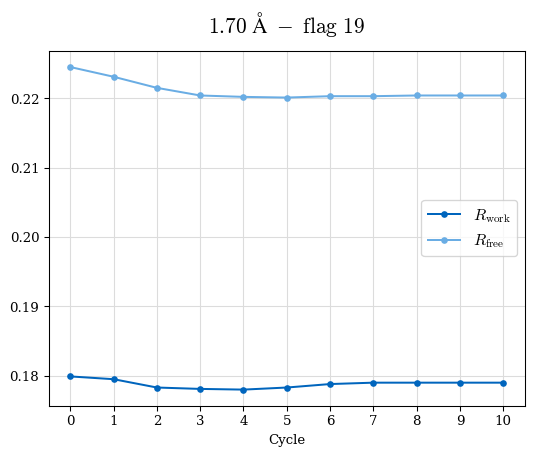
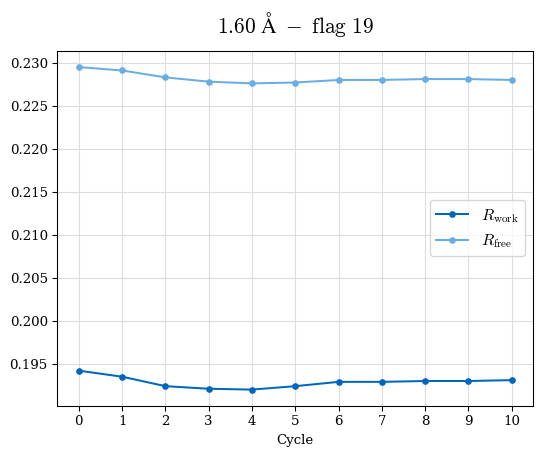
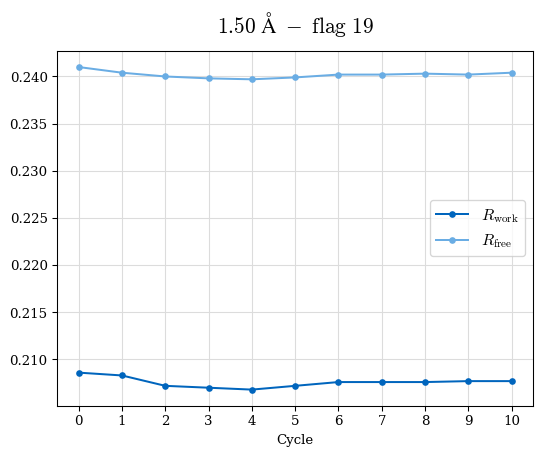
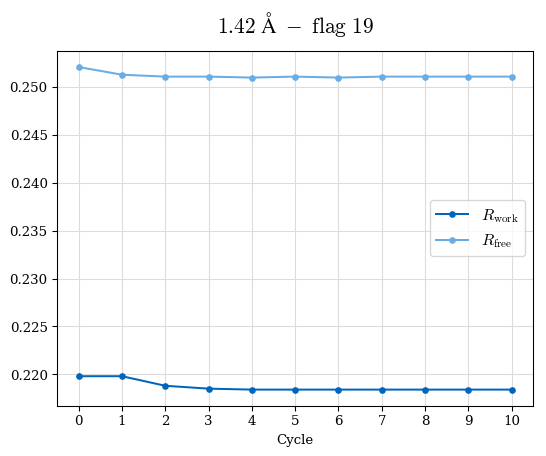
# Shell Rwork(init) Rwork(fin) Rwork(diff) Rfree(init) Rfree(fin) Rfree(diff) Rwork(StDev) Rfree(StDev) Rwork(diff,SEM) Rfree(diff,SEM) 2.00A->1.90A 0.1555 0.1566 0.0011 0.2059 0.2033 -0.0026 0.0011 0.0076 0.00015 0.00051 1.90A->1.80A 0.1582 0.1598 0.0015 0.2050 0.2029 -0.0021 0.0005 0.0062 0.00014 0.0004 1.80A->1.70A 0.1663 0.1677 0.0014 0.2091 0.2070 -0.0020 0.0007 0.0058 0.00016 0.00036 1.70A->1.60A 0.1781 0.1789 0.0008 0.2161 0.2151 -0.0010 0.0006 0.0073 0.00012 0.00019 1.60A->1.50A 0.1925 0.1925 0.0000 0.2265 0.2261 -0.0005 0.0006 0.0072 5e-05 0.00016 1.50A->1.42A 0.2071 0.2070 -0.0001 0.2382 0.2384 0.0002 0.0005 0.0066 0.0001 0.00011
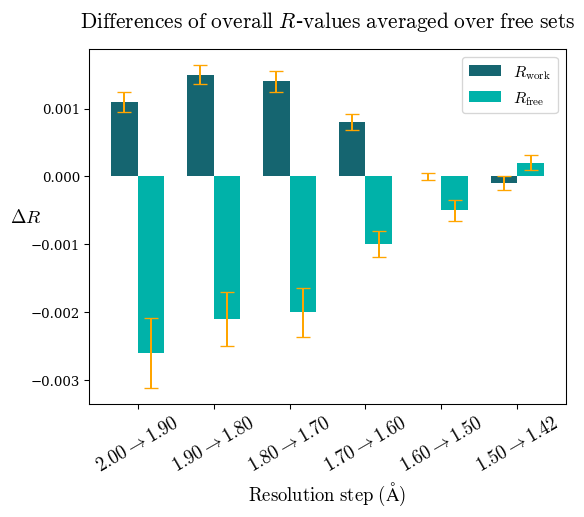
Note: For each incremental step of resolution from X->Y, the R-values were calculated at resolution X. Standard error of mean is shown in orange.