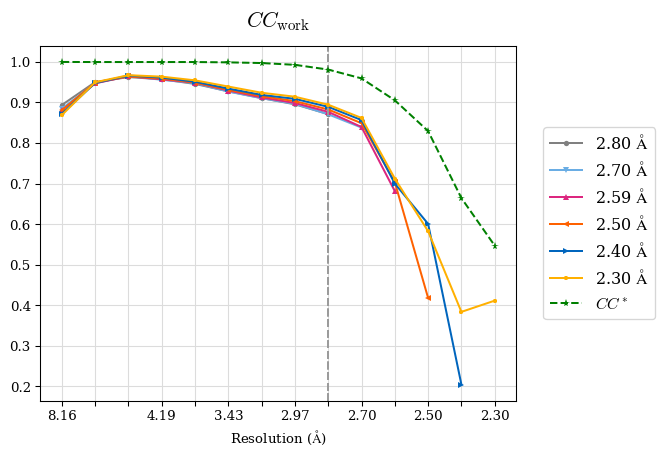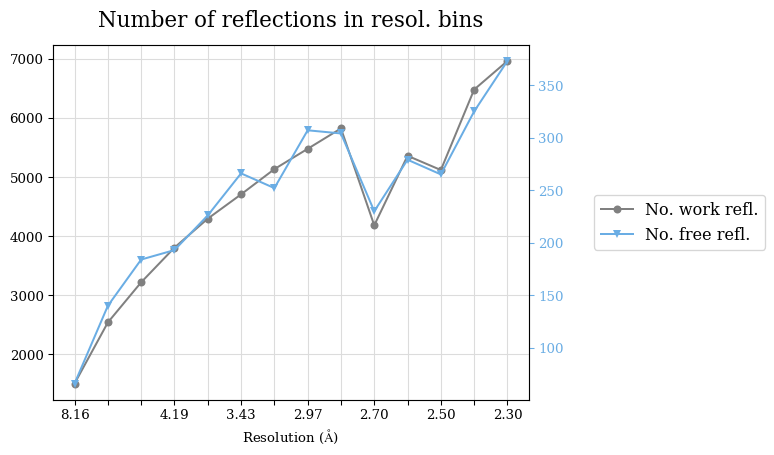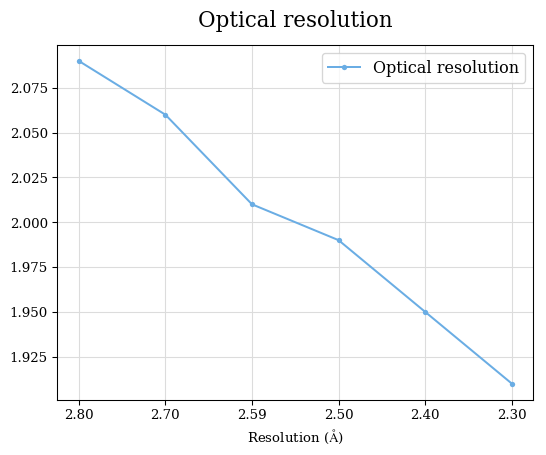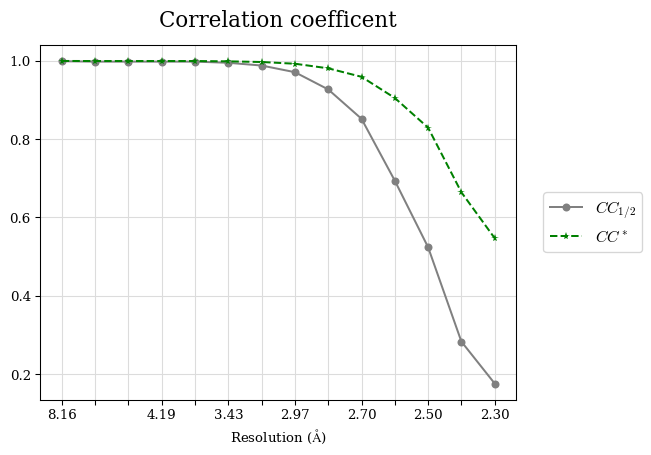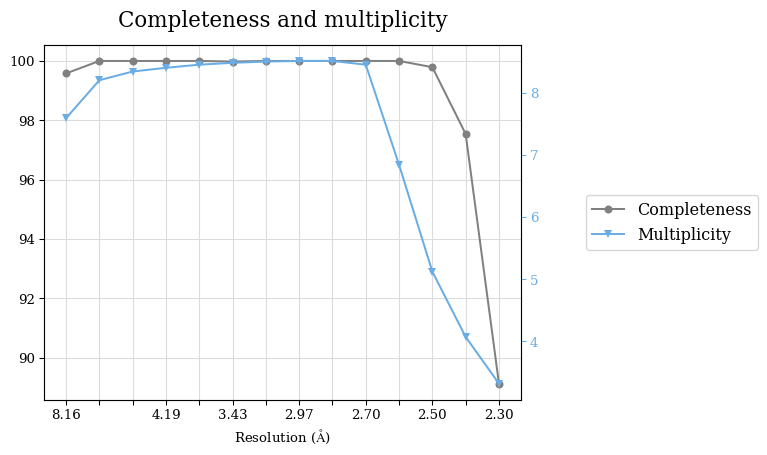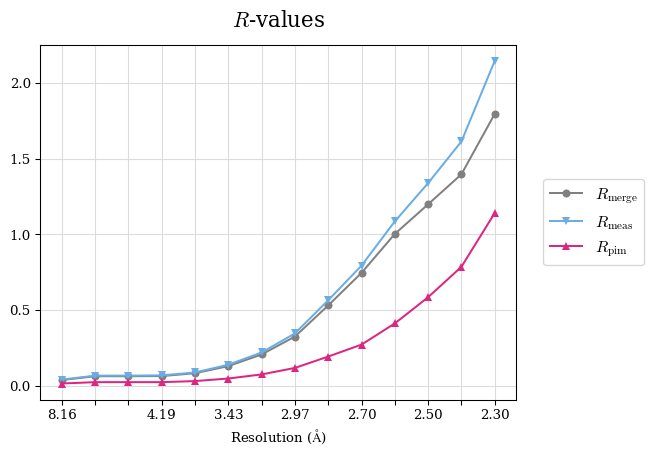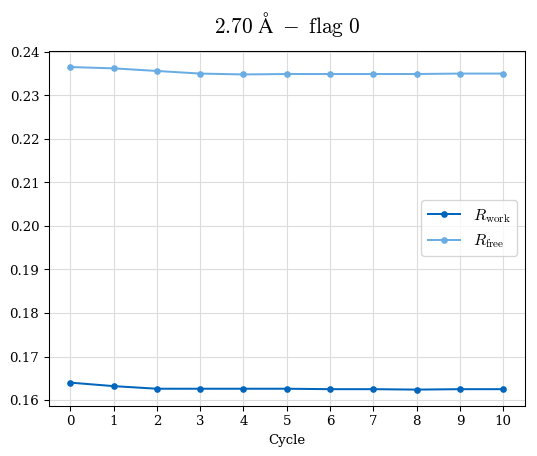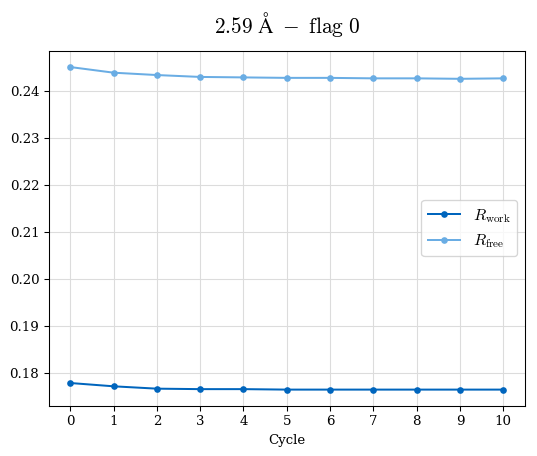
Raw data: BO_from2-80A_R-values.csv
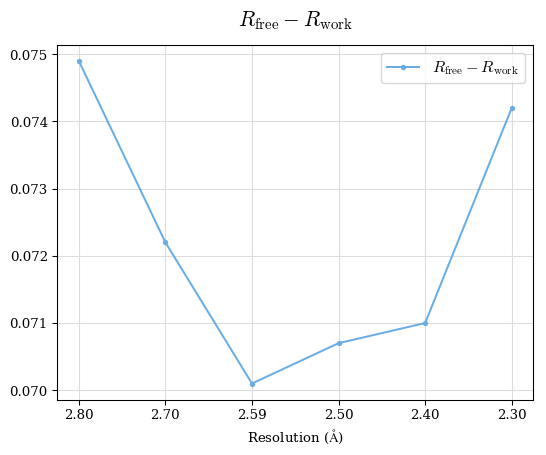
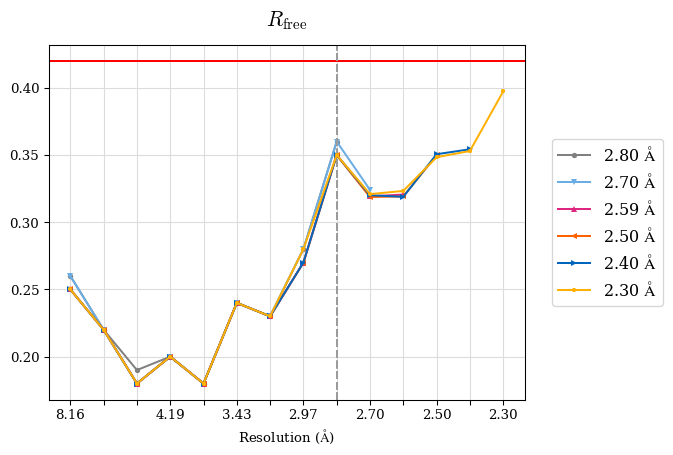
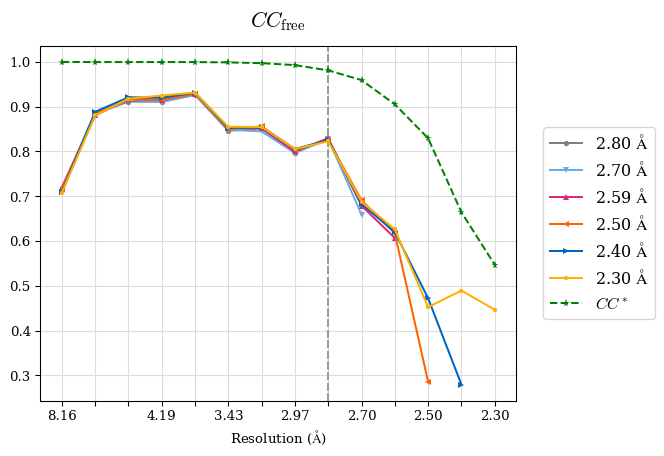
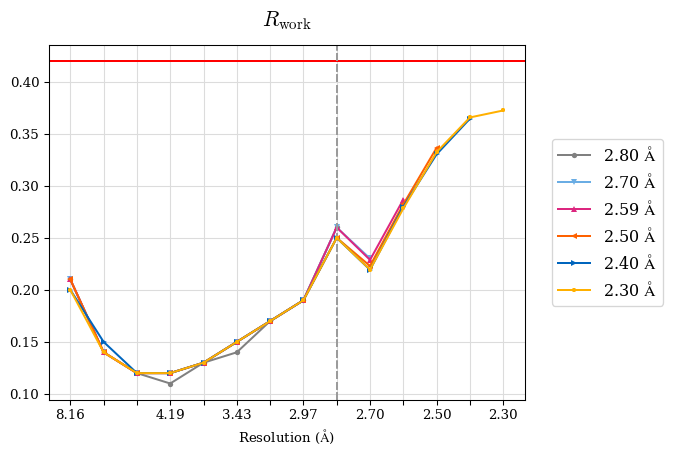
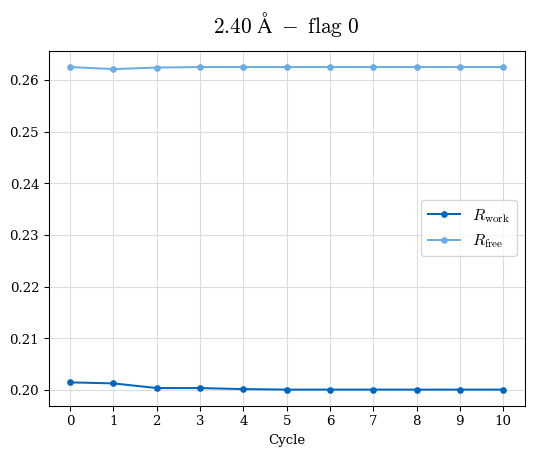
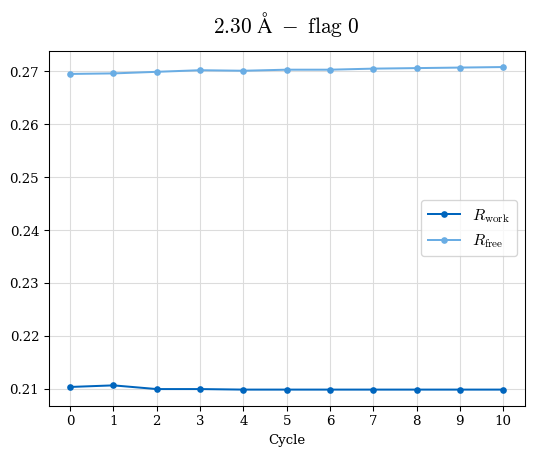
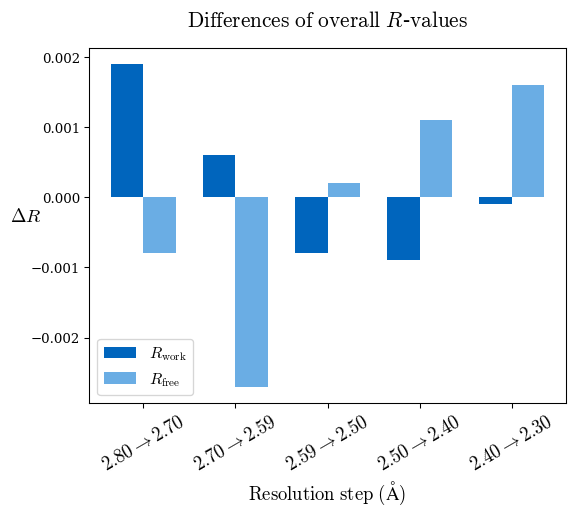
# Shell Rwork(init) Rwork(fin) Rwork(diff) Rfree(init) Rfree(fin) Rfree(diff) 2.80A->2.70A 0.1533 0.1552 0.0019 0.2282 0.2274 -0.0008 2.70A->2.59A 0.1625 0.1631 0.0006 0.2350 0.2323 -0.0027 2.59A->2.50A 0.1765 0.1757 -0.0008 0.2426 0.2428 0.0002 2.50A->2.40A 0.1882 0.1873 -0.0009 0.2514 0.2525 0.0011 2.40A->2.30A 0.2001 0.2000 -0.0001 0.2625 0.2641 0.0016
Note: For each incremental step of resolution from X->Y, the R-values were calculated at resolution X.