PAIREF webinar
23 Mar 2021, 3pm (CET)
IMPORTANT: Please install PAIREF and download data before the webinar (instructions below).
- Introduction to paired refinement. Download presentation in PDF.
- Installation of PAIREF for CCP4 and PHENIX users.
- Short tutorial. Download the archive with the data.
- Discussion.
Video recording from the webinar
Tutorial
PAIREF installation: IMPORTANT: Please install PAIREF before the webinar
Althought PAIREF has been developed to be installed easily, atypical computer setting can cause problems sometimes. Feel free to contact us via email (martin.maly@fjfi.cvut.cz) and we will help you.
PAIREF depends on the installation of the CCP4 Software Suite or PHENIX. In this webinar, we recommend to have the CCP4 Suite installed as we will be focused on that.
In GNU/Linux or macOS, just open the terminal.
In Windows, find the CCP4 console or Phenix Command Prompt in the Start menu and open it (see the screenshots below).
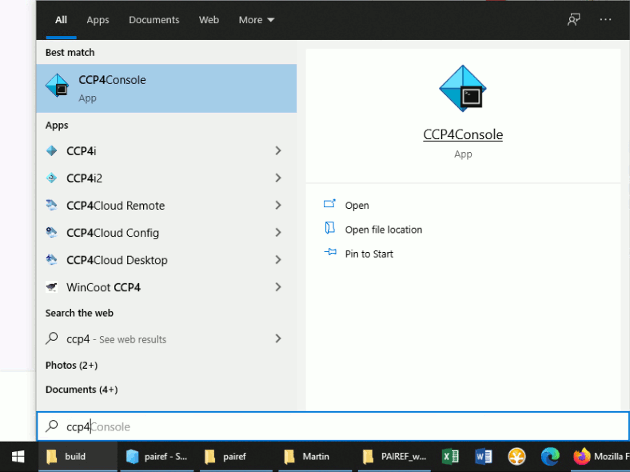
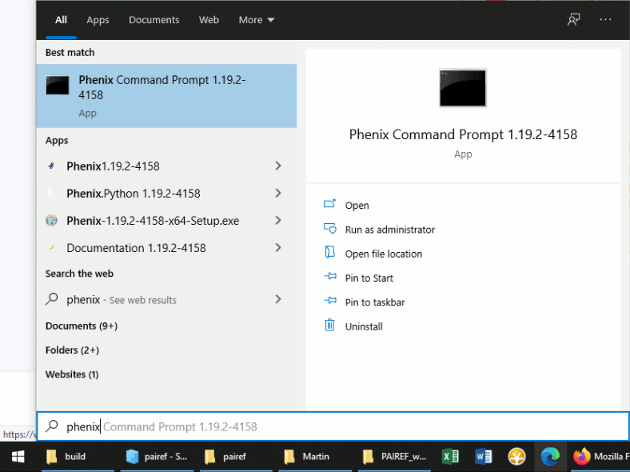
Now execute the following commands that install PAIREF into CCP4 environment on your computer:
ccp4-python -m ensurepip --user ccp4-python -m pip install pairef --no-deps --upgrade --user
If the installation was not successful, please go through the section Installation - Troubleshooting in our documentation.
In the end, please check whether the installation was successful: Try to show the help message executing the following command:
ccp4-python -m pairef --help
If the help message appears, the installation seems to be successful. Please download also the data before the webinar - the instruction follows.
Download data
In this tutorial, we will deal with the data from interferon gamma from Paralichthys olivaceus (POLI) 1), PDB entry 6F1E.
Download the archive with the data and extract it in a working folder. The archive contains:
- Structure model previously refined against data at 2.3 Å - poli67_webinar_2-3A.pdb.
- Merged diffraction data up to 1.9 Å - poli67_1-9A.mtz
- Unmerged diffraction data up to 1.9 Å - poli67_XDS_ASCII_1-9A.HKL
If you have installed PAIREF and extract the data, you are ready for the webinar. 
Open console in working directory
Open terminal (GNU/Linux, macOS) or CCP4 console (Windows) and go to the folder where your structure model and diffraction data are saved. For example, if you saved those three files to folder C:/Users/Lab/PAIREF_tutorial_poli, write cd C:/Users/Lab/PAIREF_tutorial_poli into the console and press Enter.
Run PAIREF in command line
We will add three high-resolution shells step by step: 2.3-2.10 Å, 2.1-2.0 Å and 2.0-1.9 Å. Run PAIREF:
ccp4-python -m pairef --XYZIN poli67_webinar_2-3A.pdb --HKLIN poli67_1-9A.mtz -u poli67_XDS_ASCII_1-9A.HKL -r 2.1,2.0,1.9 -w 0.06 -p poli_webinar
Note: The structure model was refined using X-ray weight 0.06, we have to keep this setting to gain unbiased results.
Graphical user interface
PAIREF provides also a graphical user interface (GUI) for intuitive setting of arguments without making up a command. However, it depends on the installation of CCP4 suite. For launching the PAIREF GUI, execute the following command:
ccp4-python -m pairef --gui
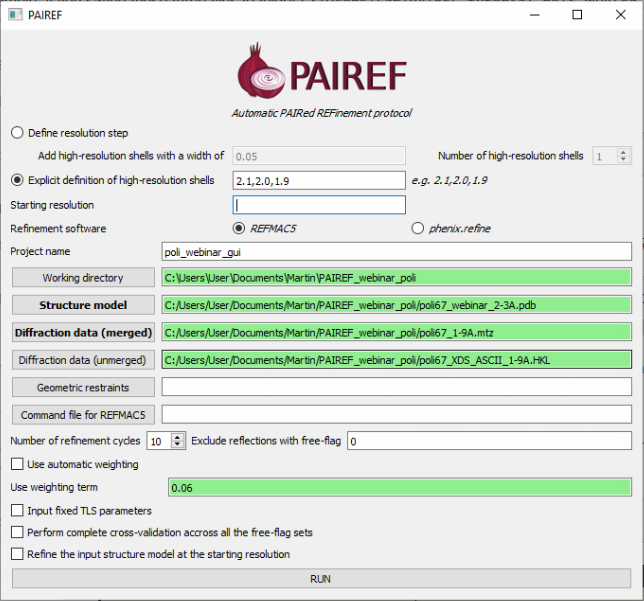
Interpretation of results
Follow the note ------> RESULTS AND THE CURRENT STATUS OF CALCULATIONS ARE LISTED IN A HTML LOG FILE in the program output and open the stated file in your preferred web browser (e.g. Firefox). The results should look similar to ours: https://pairef.fjfi.cvut.cz/docs/pairef_poli_webinar/PAIREF_poli_webinar.html.
The first thing that should be checked is whether the refinements have converged. Scroll at the very bottom of page, here you can see plots of Rwork and Rfree vs. refinement cycle. We can conclude that all the refinements have converged.
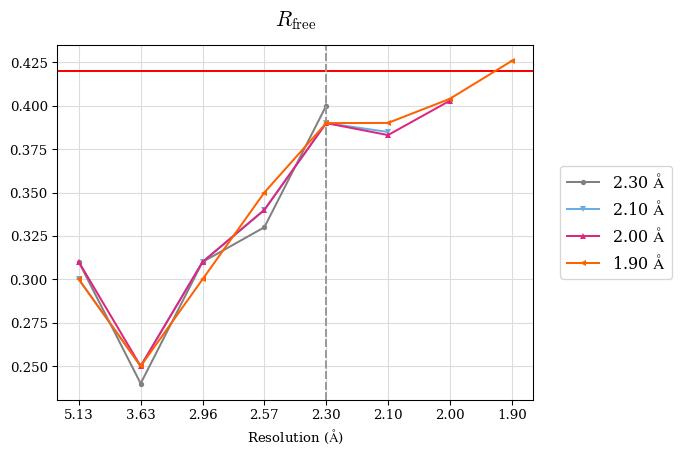
Rfree decreased after the addition of shells 2.3-2.10 Å and 2.1-2.0 Å:
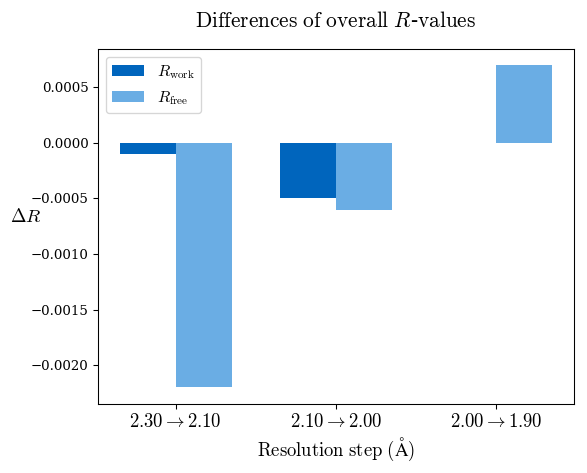
Since a perfect model gives an R-value of 0.42 against random data (i.e. pure noise) – assuming non-tNCS (translational non-crystallographic symmetry) data from a non-twinned crystal 2) – a higher R-value in the (current) high-resolution shell indicates either the involvement of high-resolution data without information content (the data are even worse than noise), or poor quality of the model, or the presence of tNCS. This is indicated for the shell 2.0-1.9 Å.
CC* a model-independent measure of noise is in the diffraction data. CC* is higher than CCwork in whole resolution range (except the shell 2.0-1.9 Å where CC* is undefined due to negative CC1/2. Thus, the overfitting was not indicated. To access overfitting, it is not needed to test set, so the comparison of CC* with CCwork is much better then with CCfree as CCwork is calculated on more data.
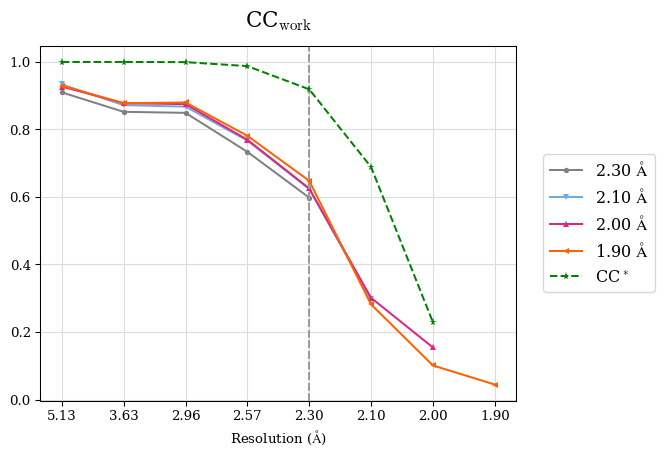
We can conclude that the high-resolution cutoff can be set to 2.0 Å.
Merging statistics:
#shell d_max d_min #obs #uniq mult. %comp <I> <I/sI> r_mrg r_meas r_pim cc1/2 cc_ano cc* 01 47.05 5.13 22367 1948 11.48 99.44 494.1 38.8 0.042 0.044 0.013 0.999 -0.214 0.9997 02 5.13 3.63 42232 3343 12.63 99.79 203.3 31.7 0.064 0.067 0.019 0.999 -0.163 0.9997 03 3.63 2.97 57138 4246 13.46 99.95 56.9 14.3 0.151 0.157 0.042 0.997 -0.166 0.9992 04 2.97 2.57 65007 5014 12.97 99.70 11.3 4.0 0.622 0.648 0.178 0.951 -0.025 0.9874 05 2.57 2.30 73887 5623 13.14 99.89 3.7 1.3 1.839 1.914 0.523 0.730 -0.014 0.9187 06 2.30 2.10 83129 6181 13.45 99.92 1.2 0.4 5.812 6.041 1.633 0.311 0.001 0.6888 07 2.10 2.00 49772 4021 12.38 98.51 0.3 0.1 16.989 17.721 4.963 0.027 0.008 0.2293 08 2.00 1.90 35920 4046 8.88 81.18 0.1 0.0 41.435 43.993 14.417 -0.132 -0.016 N/A
Contact
In case of any questions or problems, please do not hesitate and write us: martin.maly@fjfi.cvut.cz.
Further reading
- More information about PAIREF settings and possibilities are explained in the documentation.
- Linking crystallographic model and data quality. P.A. Karplus & K. Diederichs (2012) Science 336:1030–3
- Assessing and maximizing data quality in macromolecular crystallography. P.A. Karplus & K. Diederichs (2015) Cur. Op. in Str. Biology 34:60–68
- Better models by discarding data? P.A. Karplus & K. Diederichs (2013) Acta Cryst. D59:1215–1222
